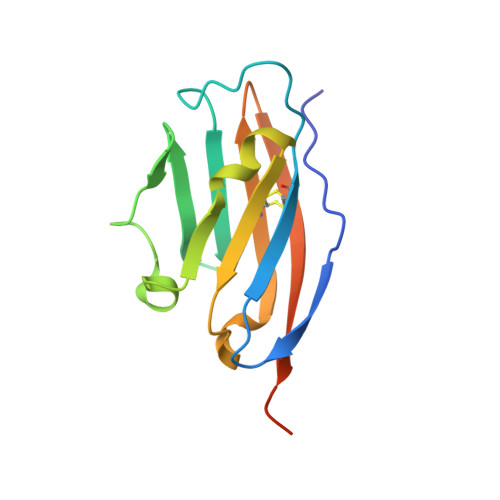
Structural Characterization of a Macrocyclic Peptide Modulator of the PD-1/PD-L1 Immune Checkpoint Axis.
Zyla, E., Musielak, B., Holak, T.A., Dubin, G.(2021) Molecules 26
- PubMed: 34443436 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/molecules26164848
- Primary Citation Related Structures:
7OUN - PubMed Abstract:
The clinical success of PD-1/PD-L1 immune checkpoint targeting antibodies in cancer is followed by efforts to develop small molecule inhibitors with better penetration into solid tumors and more favorable pharmacokinetics. Here we report the crystal structure of a macrocyclic peptide inhibitor (peptide 104) in complex with PD-L1. Our structure shows no indication of an unusual bifurcated binding mode demonstrated earlier for another peptide of the same family (peptide 101). The binding mode relies on extensive hydrophobic interactions at the center of the binding surface and an electrostatic patch at the side. An interesting sulfur/π interaction supports the macrocycle-receptor binding. Overall, our results allow a better understanding of forces guiding macrocycle affinity for PD-L1, providing a rationale for future structure-based inhibitor design and rational optimization.
- Malopolska Centre of Biotechnology, Jagiellonian University, Gronostajowa 7A, 30-387 Krakow, Poland.
Organizational Affiliation: