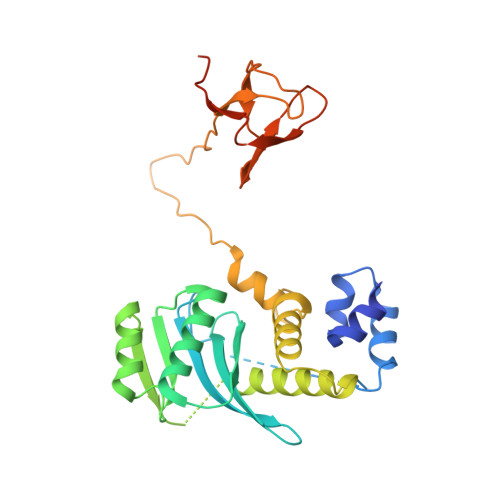
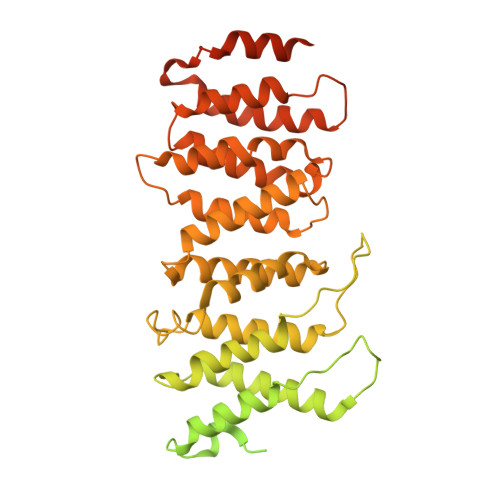
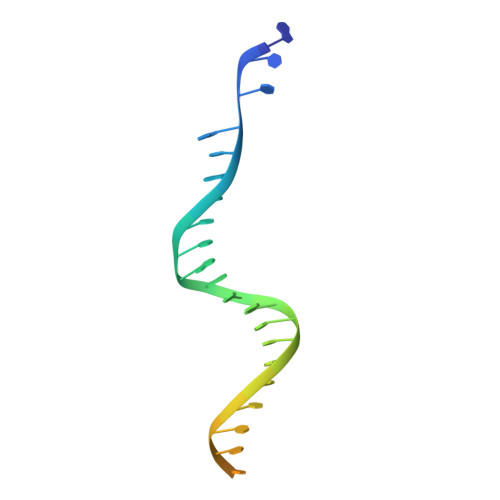
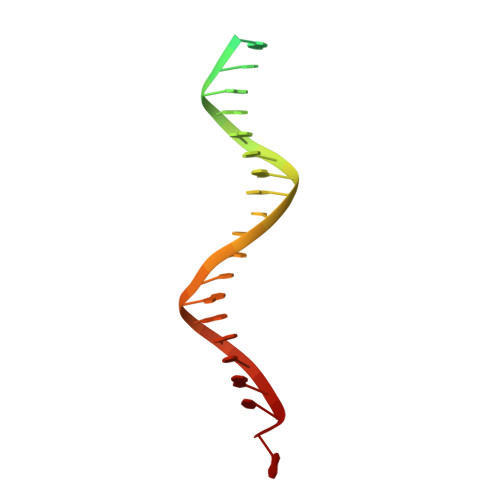
Structural basis for the inhibition of HTLV-1 integration inferred from cryo-EM deltaretroviral intasome structures.
Barski, M.S., Vanzo, T., Zhao, X.Z., Smith, S.J., Ballandras-Colas, A., Cronin, N.B., Pye, V.E., Hughes, S.H., Burke Jr., T.R., Cherepanov, P., Maertens, G.N.(2021) Nat Commun 12: 4996-4996
- PubMed: 34404793 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-25284-1
- Primary Citation Related Structures:
7OUF, 7OUG, 7OUH - PubMed Abstract:
Between 10 and 20 million people worldwide are infected with the human T-cell lymphotropic virus type 1 (HTLV-1). Despite causing life-threatening pathologies there is no therapeutic regimen for this deltaretrovirus. Here, we screened a library of integrase strand transfer inhibitor (INSTI) candidates built around several chemical scaffolds to determine their effectiveness in limiting HTLV-1 infection. Naphthyridines with substituents in position 6 emerged as the most potent compounds against HTLV-1, with XZ450 having highest efficacy in vitro. Using single-particle cryo-electron microscopy we visualised XZ450 as well as the clinical HIV-1 INSTIs raltegravir and bictegravir bound to the active site of the deltaretroviral intasome. The structures reveal subtle differences in the coordination environment of the Mg 2+ ion pair involved in the interaction with the INSTIs. Our results elucidate the binding of INSTIs to the HTLV-1 intasome and support their use for pre-exposure prophylaxis and possibly future treatment of HTLV-1 infection.
- Imperial College London, St. Mary's Hospital, Department of Infectious Disease, Section of Virology, Norfolk Place, London, UK.
Organizational Affiliation: