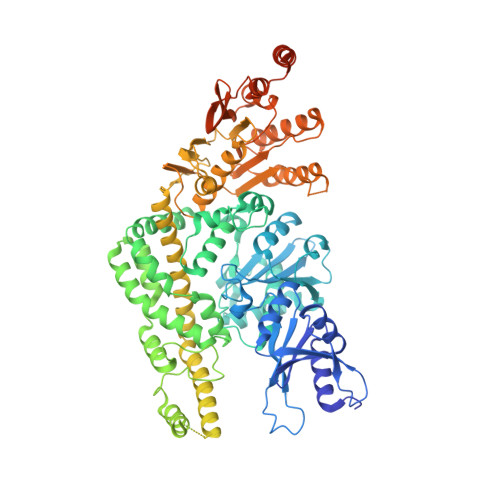
Cryogenic electron microscopy structures reveal how ATP and DNA binding in MutS coordinates sequential steps of DNA mismatch repair.
Borsellini, A., Kunetsky, V., Friedhoff, P., Lamers, M.H.(2022) Nat Struct Mol Biol 29: 59-66
- PubMed: 35013597 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00707-1
- Primary Citation Related Structures:
7OTO, 7OU0, 7OU2, 7OU4 - PubMed Abstract:
DNA mismatch repair detects and corrects mismatches introduced during DNA replication. The protein MutS scans for mismatches and coordinates the repair cascade. During this process, MutS undergoes multiple conformational changes in response to ATP binding, hydrolysis and release, but how ATP induces the various MutS conformations is incompletely understood. Here we present four cryogenic electron microscopy structures of Escherichia coli MutS at sequential stages of the ATP hydrolysis cycle that reveal how ATP binding and hydrolysis induce closing and opening of the MutS dimer, respectively. Biophysical analysis demonstrates how DNA binding modulates the ATPase cycle by prevention of hydrolysis during scanning and mismatch binding, while preventing ADP release in the sliding clamp state. Nucleotide release is achieved when MutS encounters single-stranded DNA that is produced during removal of the daughter strand. The combination of ATP binding and hydrolysis and its modulation by DNA enables MutS to adopt the different conformations needed to coordinate the sequential steps of the mismatch repair cascade.
- Department of Cell and Chemical Biology, Leiden University Medical Center, Leiden, the Netherlands.
Organizational Affiliation: