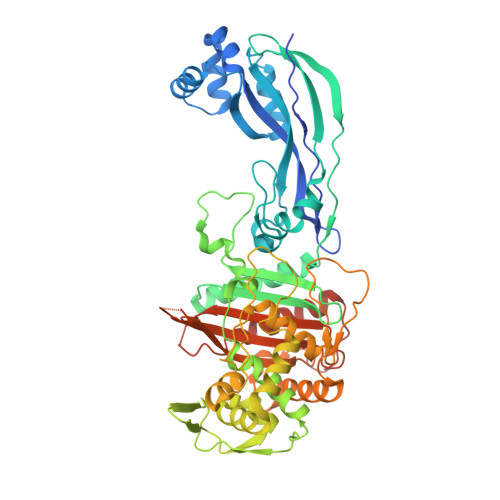
Interaction Mode of the Novel Monobactam AIC499 Targeting Penicillin Binding Protein 3 of Gram-Negative Bacteria.
Freischem, S., Grimm, I., Lopez-Perez, A., Willbold, D., Klenke, B., Vuong, C., Dingley, A.J., Weiergraber, O.H.(2021) Biomolecules 11
- PubMed: 34356681 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/biom11071057
- Primary Citation Related Structures:
7ONK, 7ONN, 7ONO, 7ONW, 7ONX, 7ONY, 7ONZ - PubMed Abstract:
Novel antimicrobial strategies are urgently required because of the rising threat of multi drug resistant bacterial strains and the infections caused by them. Among the available target structures, the so-called penicillin binding proteins are of particular interest, owing to their good accessibility in the periplasmic space, and the lack of homologous proteins in humans, reducing the risk of side effects of potential drugs. In this report, we focus on the interaction of the innovative β-lactam antibiotic AIC499 with penicillin binding protein 3 (PBP3) from Escherichia coli and Pseudomonas aeruginosa . This recently developed monobactam displays broad antimicrobial activity, against Gram-negative strains, and improved resistance to most classes of β-lactamases. By analyzing crystal structures of the respective complexes, we were able to explore the binding mode of AIC499 to its target proteins. In addition, the apo structures determined for PBP3, from P. aeruginosa and the catalytic transpeptidase domain of the E. coli orthologue, provide new insights into the dynamics of these proteins and the impact of drug binding.
- Institute of Biological Information Processing (IBI-7: Structural Biochemistry) and Jülich Centre for Structural Biology (JuStruct), Forschungszentrum Jülich, 52425 Jülich, Germany.
Organizational Affiliation: