Deconstruction of Dual-Site Tankyrase Inhibitors Provides Insights into Binding Energetics and Suggests Critical Hotspots for Ligand Optimization.
Sowa, S.T., Kucukdisli, M., Mostinski, Y., Schaller, D.A., Vinagreiro, C.S., Cirillo, D., Bosetti, C., Brinch, S.A., van Laar, K., Wegert, A., Leenders, R.G.G., Krauss, S., Waaler, J., Volkamer, A., Lehtio, L., Nazare, M.(2025) J Med Chem 68: 7263-7279
- PubMed: 40134122 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.4c02845
- Primary Citation Related Structures:
7OJO, 8B6M - PubMed Abstract:
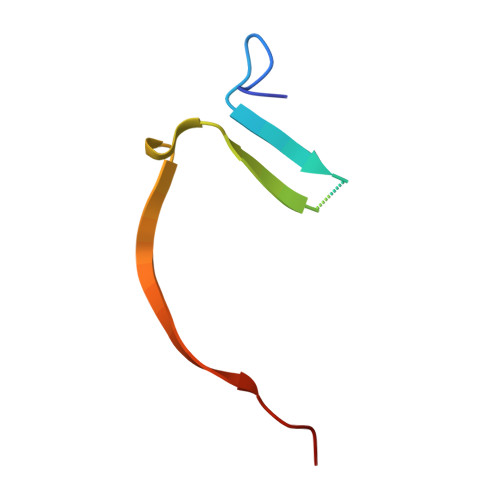
Designing inhibitors is a complex task that requires a deep understanding of protein-ligand interactions and their dynamics. Ligands often interact with multiple binding subsites, with noncovalent interactions affecting binding affinity. Conformational changes and plasticity of both, the ligand and the protein influence binding energetics. We investigated the tankyrase ADP-ribosyltransferase as a promising drug target regulating many cellular pathways. Despite the existence of diverse tankyrase inhibitors, their binding energetics and contributions of flexible cryptic subpockets to binding affinity remain elusive. To examine these aspects, we deconstructed inhibitors to key fragments, dissected their energetic contribution to the affinity, and determined their binding mode by X-ray crystallography. Varying ligand efficiencies of the deconstructed, pocket-binding fragments revealed the cryptic nature of subpockets. These insights enabled us to redesign inhibitors with novel linkers, the observed key area for optimization, increasing the potency in enzymatic and cell-based assays by 7.5-fold and 6.2-fold compared to the parent ligand.
- Faculty for Biochemistry and Molecular Medicine & Biocenter Oulu, University of Oulu, Aapistie 7, 90220 Oulu, Finland.
Organizational Affiliation: