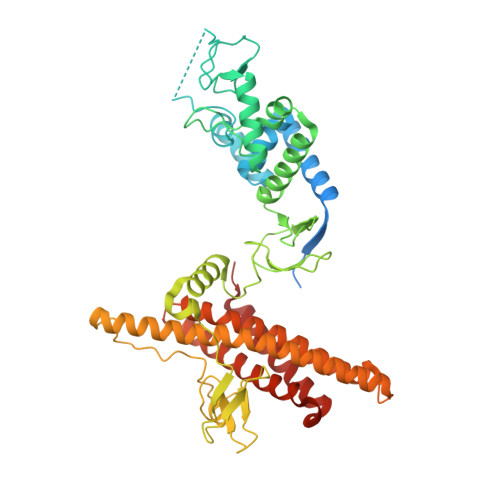
CUL5-ARIH2 E3-E3 ubiquitin ligase structure reveals cullin-specific NEDD8 activation.
Kostrhon, S., Prabu, J.R., Baek, K., Horn-Ghetko, D., von Gronau, S., Klugel, M., Basquin, J., Alpi, A.F., Schulman, B.A.(2021) Nat Chem Biol 17: 1075-1083
- PubMed: 34518685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-021-00858-8
- Primary Citation Related Structures:
7OD1, 7ONI - PubMed Abstract:
An emerging mechanism of ubiquitylation involves partnering of two distinct E3 ligases. In the best-characterized E3-E3 pathways, ARIH-family RING-between-RING (RBR) E3s ligate ubiquitin to substrates of neddylated cullin-RING E3s. The E3 ARIH2 has been implicated in ubiquitylation of substrates of neddylated CUL5-RBX2-based E3s, including APOBEC3-family substrates of the host E3 hijacked by HIV-1 virion infectivity factor (Vif). However, the structural mechanisms remained elusive. Here structural and biochemical analyses reveal distinctive ARIH2 autoinhibition, and activation on assembly with neddylated CUL5-RBX2. Comparison to structures of E3-E3 assemblies comprising ARIH1 and neddylated CUL1-RBX1-based E3s shows cullin-specific regulation by NEDD8. Whereas CUL1-linked NEDD8 directly recruits ARIH1, CUL5-linked NEDD8 does not bind ARIH2. Instead, the data reveal an allosteric mechanism. NEDD8 uniquely contacts covalently linked CUL5, and elicits structural rearrangements that unveil cryptic ARIH2-binding sites. The data reveal how a ubiquitin-like protein induces protein-protein interactions indirectly, through allostery. Allosteric specificity of ubiquitin-like protein modifications may offer opportunities for therapeutic targeting.
- Department of Molecular Machines and Signaling, Max Planck Institute of Biochemistry, Martinsried, Germany.
Organizational Affiliation: