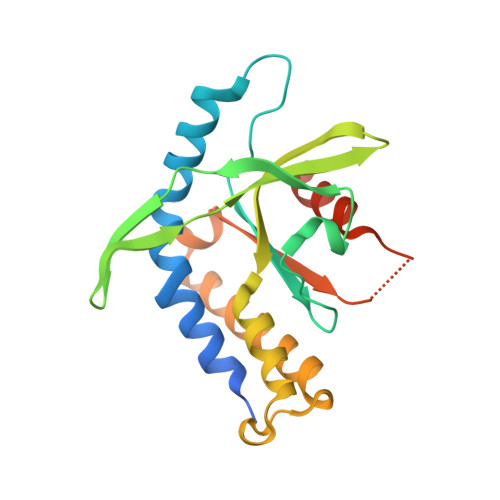
Enzymatic Synthesis of 3'-5', 3'-5' Cyclic Dinucleotides, Their Binding Properties to the Stimulator of Interferon Genes Adaptor Protein, and Structure/Activity Correlations
Novotna, B., Hola, L., Stas, M., Gutten, O., Smola, M., Zavrel, M., Vavrina, Z., Budesinsky, M., Liboska, R., Chevrier, F., Dobias, J., Boura, E., Rulisek, L., Birkus, G.(2021) Biochemistry