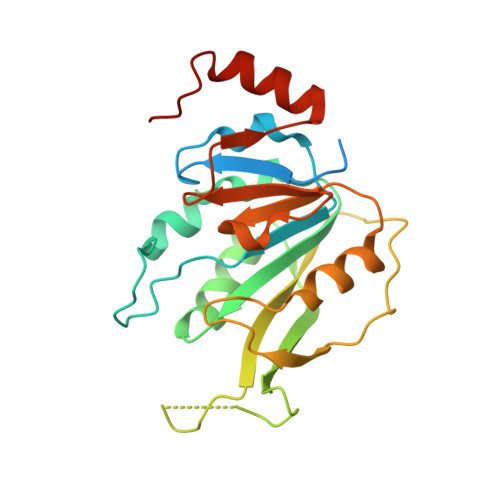
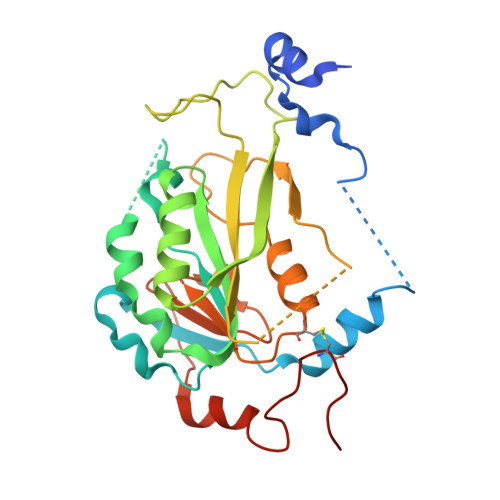
1,4,9-Triazaspiro[5.5]undecan-2-one Derivatives as Potent and Selective METTL3 Inhibitors.
Dolbois, A., Bedi, R.K., Bochenkova, E., Muller, A., Moroz-Omori, E.V., Huang, D., Caflisch, A.(2021) J Med Chem 64: 12738-12760
- PubMed: 34431664 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c00773
- Primary Citation Related Structures:
7O08, 7O09, 7O0L, 7O29, 7O2E, 7O2F - PubMed Abstract:
N 6 -methyladenosine (m 6 A) is the most frequent of the 160 RNA modifications reported so far. Accumulating evidence suggests that the METTL3/METTL14 protein complex, part of the m 6 A regulation machinery, is a key player in a variety of diseases including several types of cancer, type 2 diabetes, and viral infections. Here we report on a protein crystallography-based medicinal chemistry optimization of a METTL3 hit compound that has resulted in a 1400-fold potency improvement (IC 50 of 5 nM for the lead compound 22 ( UZH2 ) in a time-resolved Förster resonance energy transfer (TR-FRET) assay). The series has favorable ADME properties as physicochemical characteristics were taken into account during hit optimization. UZH2 shows target engagement in cells and is able to reduce the m 6 A/A level of polyadenylated RNA in MOLM-13 (acute myeloid leukemia) and PC-3 (prostate cancer) cell lines.
- Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, Zurich CH-8057, Switzerland.
Organizational Affiliation: