Conformational plasticity of the HIV-1 gp41 immunodominant region is recognized by multiple non-neutralizing antibodies.
Cook, J.D., Khondker, A., Lee, J.E.(2022) Commun Biol 5: 291-291
- PubMed: 35361878
- DOI: https://doi.org/10.1038/s42003-022-03235-w
- Primary Citation Related Structures:
7N04, 7N05, 7N07, 7N08 - PubMed Abstract:
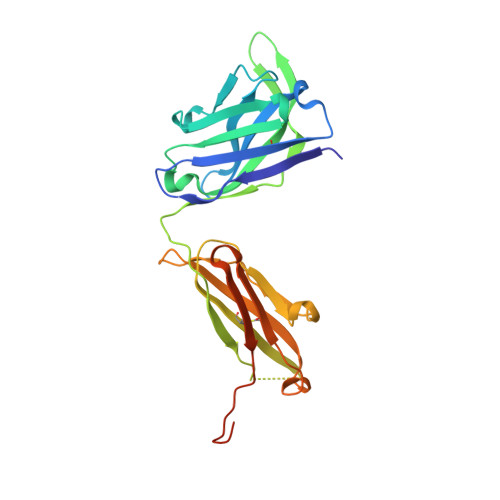
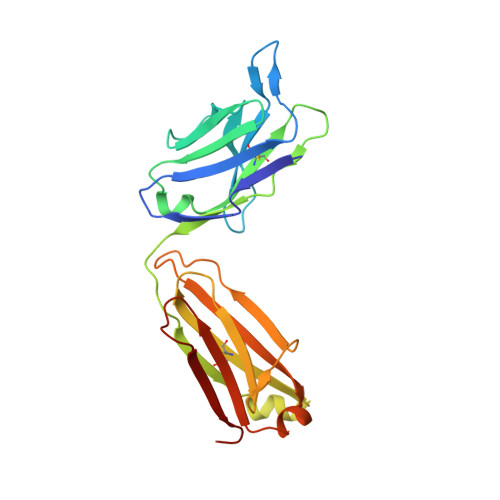
The early humoral immune response to acute HIV-1 infection is largely non-neutralizing. The principal target of these antibodies is the primary immunodominant region (PID) on the gp41 fusion protein. The PID is a highly conserved 15-residue region displayed on the surface of HIV-1 virions. In this study, we analyzed the humoral determinants of HIV-1 gp41 PID binding using biophysical, structural, and computational methods. In complex with a patient-derived near-germline antibody fragment, the PID motif adopts an elongated random coil, whereas the PID bound to affinity-matured Fab adopts a strand-turn-helix conformation. Molecular dynamics simulations showed that the PID is structurally plastic suggesting that the PID can form an ensemble of structural states recognized by various non-neutralizing antibodies, facilitating HIV-1 immunodominance observed in acute and chronic HIV-1 infections. An improved understanding of how the HIV-1 gp41 PID misdirects the early humoral response should guide the development of an effective HIV-1 vaccine.
- Department of Laboratory Medicine and Pathobiology, Temerty Faculty of Medicine, University of Toronto, Toronto, ON, M5S 1A8, Canada.
Organizational Affiliation: