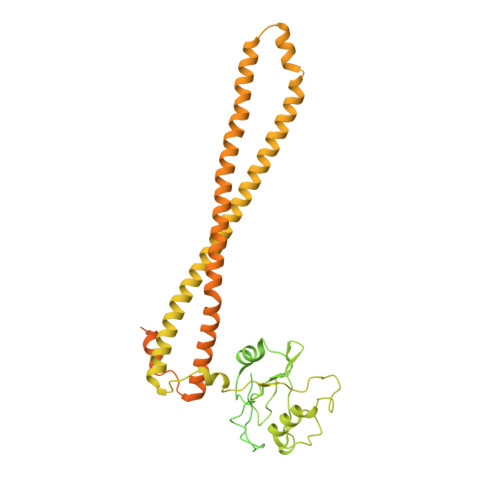
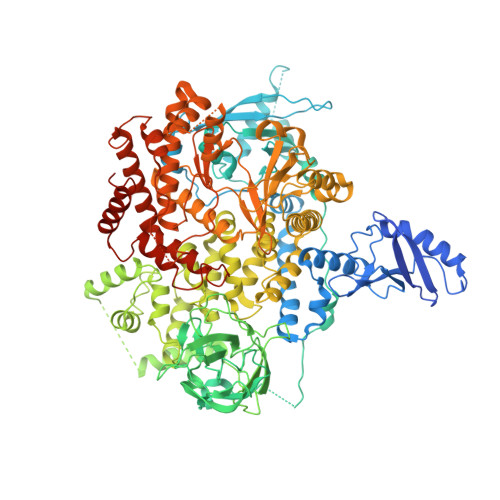
Cryo-EM structures of PI3K alpha reveal conformational changes during inhibition and activation.
Liu, X., Yang, S., Hart, J.R., Xu, Y., Zou, X., Zhang, H., Zhou, Q., Xia, T., Zhang, Y., Yang, D., Wang, M.W., Vogt, P.K.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 34725156
- DOI: https://doi.org/10.1073/pnas.2109327118
- Primary Citation Related Structures:
7MYN, 7MYO - PubMed Abstract:
Phosphoinositide 3-kinases (PI3Ks) are lipid kinases essential for growth and metabolism. Their aberrant activation is associated with many types of cancers. Here we used single-particle cryoelectron microscopy (cryo-EM) to determine three distinct conformations of full-length PI3Kα (p110α-p85α): the unliganded heterodimer PI3Kα, PI3Kα bound to the p110α-specific inhibitor BYL-719, and PI3Kα exposed to an activating phosphopeptide. The cryo-EM structures of unbound and of BYL-719-bound PI3Kα are in general accord with published crystal structures. Local deviations are presented and discussed. BYL-719 stabilizes the structure of PI3Kα, but three regions of low-resolution extra density remain and are provisionally assigned to the cSH2, BH, and SH3 domains of p85. One of the extra density regions is in contact with the kinase domain blocking access to the catalytic site. This conformational change indicates that the effects of BYL-719 on PI3Kα activity extend beyond competition with adenosine triphosphate (ATP). In unliganded PI3Kα, the DFG motif occurs in the "in" and "out" positions. In BYL-719-bound PI3Kα, only the DFG-in position, corresponding to the active conformation of the kinase, was observed. The phosphopeptide-bound structure of PI3Kα is composed of a stable core resolved at 3.8 Å. It contains all p110α domains except the adaptor-binding domain (ABD). The p85α domains, linked to the core through the ABD, are no longer resolved, implying that the phosphopeptide activates PI3Kα by fully releasing the niSH2 domain from binding to p110α. The structures presented here show the basal form of the full-length PI3Kα dimer and document conformational changes related to the activated and inhibited states.
- School of Pharmacy, Fudan University, Shanghai 201203, China.
Organizational Affiliation: