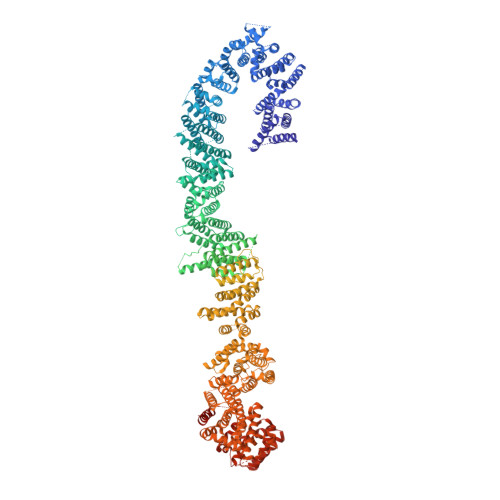
The cryo-EM structure of the human neurofibromin dimer reveals the molecular basis for neurofibromatosis type 1.
Lupton, C.J., Bayly-Jones, C., D'Andrea, L., Huang, C., Schittenhelm, R.B., Venugopal, H., Whisstock, J.C., Halls, M.L., Ellisdon, A.M.(2021) Nat Struct Mol Biol 28: 982-988
- PubMed: 34887559 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-021-00687-2
- Primary Citation Related Structures:
7MOC, 7MP5, 7MP6 - PubMed Abstract:
Neurofibromin (NF1) mutations cause neurofibromatosis type 1 and drive numerous cancers, including breast and brain tumors. NF1 inhibits cellular proliferation through its guanosine triphosphatase-activating protein (GAP) activity against rat sarcoma (RAS). In the present study, cryo-electron microscope studies reveal that the human ~640-kDa NF1 homodimer features a gigantic 30 × 10 nm array of α-helices that form a core lemniscate-shaped scaffold. Three-dimensional variability analysis captured the catalytic GAP-related domain and lipid-binding SEC-PH domains positioned against the core scaffold in a closed, autoinhibited conformation. We postulate that interaction with the plasma membrane may release the closed conformation to promote RAS inactivation. Our structural data further allow us to map the location of disease-associated NF1 variants and provide a long-sought-after structural explanation for the extreme susceptibility of the molecule to loss-of-function mutations. Collectively these findings present potential new routes for therapeutic modulation of the RAS pathway.
- Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: