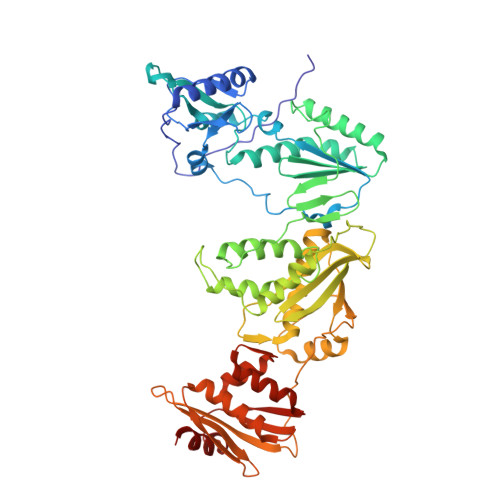
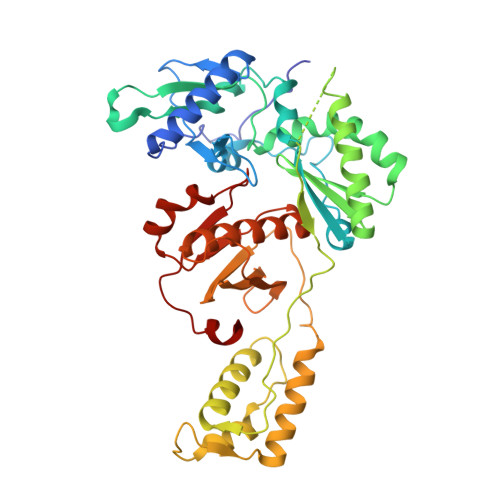
HIV-1 gp120 Antagonists Also Inhibit HIV-1 Reverse Transcriptase by Bridging the NNRTI and NRTI Sites.
Losada, N., Ruiz, F.X., Curreli, F., Gruber, K., Pilch, A., Das, K., Debnath, A.K., Arnold, E.(2021) J Med Chem 64: 16530-16540
- PubMed: 34735153
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01104
- Primary Citation Related Structures:
7LPW, 7LPX, 7LQU - PubMed Abstract:
HIV-1 infection is typically treated using ≥2 drugs, including at least one HIV-1 reverse transcriptase (RT) inhibitor. Drugs targeting RT comprise nucleos(t)ide RT inhibitors (NRTIs) and non-nucleoside RT inhibitors (NNRTIs). NRTI-triphosphates bind at the polymerase active site and, following incorporation, inhibit DNA elongation. NNRTIs bind at an allosteric pocket ∼10 Å away from the polymerase active site. This study focuses on compounds ("NBD derivatives") originally developed to bind to HIV-1 gp120, some of which inhibit RT. We have determined crystal structures of three NBD compounds in complex with HIV-1 RT, correlating with RT enzyme inhibition and antiviral activity, to develop structure-activity relationships. Intriguingly, these compounds bridge the dNTP and NNRTI-binding sites and inhibit the polymerase activity of RT in the enzymatic assays (IC 50 < 5 μM). Two of the lead compounds, NBD-14189 and NBD-14270, show potent antiviral activity (EC 50 < 200 nM), and NBD-14270 shows low cytotoxicity (CC 50 > 100 μM).
- Center for Advanced Biotechnology and Medicine, Rutgers University, Piscataway, New Jersey 08854, United States.
Organizational Affiliation: