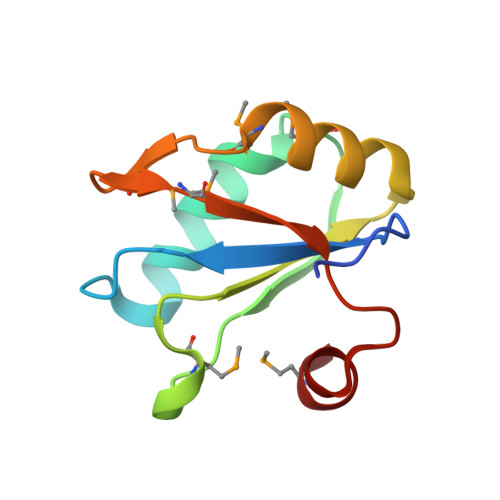
Affinity and Structural Analysis of the U1A RNA Recognition Motif with Engineered Methionines to Improve Experimental Phasing
Srivastava, K.Y., Bonn-Breach, R.B., Chavali, S.S., Lippa, G.M., Jenkins, J.L., Wedekind, J.E.(2021) Crystals (Basel) 11: 273