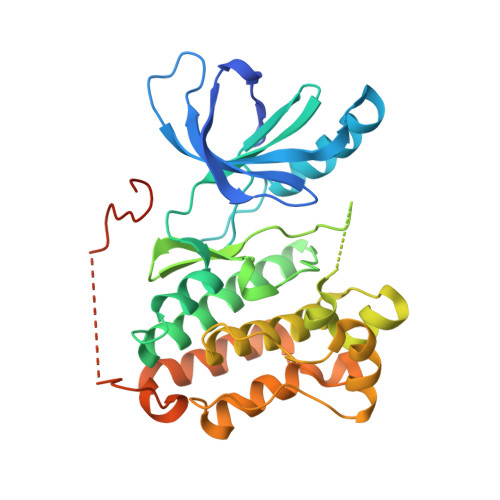
Molecular basis for cooperative binding and synergy of ATP-site and allosteric EGFR inhibitors.
Beyett, T.S., To, C., Heppner, D.E., Rana, J.K., Schmoker, A.M., Jang, J., De Clercq, D.J.H., Gomez, G., Scott, D.A., Gray, N.S., Janne, P.A., Eck, M.J.(2022) Nat Commun 13: 2530-2530
- PubMed: 35534503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-30258-y
- Primary Citation Related Structures:
6XL4, 7LG8 - PubMed Abstract:
Lung cancer is frequently caused by activating mutations in the epidermal growth factor receptor (EGFR). Allosteric EGFR inhibitors offer promise as the next generation of therapeutics, as they are unaffected by common ATP-site resistance mutations and synergize with the drug osimertinib. Here, we examine combinations of ATP-competitive and allosteric inhibitors to better understand the molecular basis for synergy. We identify a subset of irreversible EGFR inhibitors that display positive binding cooperativity and synergy with the allosteric inhibitor JBJ-04-125-02 in several EGFR variants. Structural analysis of these complexes reveals conformational changes occur mainly in the phosphate-binding loop (P-loop). Mutation of F723 in the P-loop reduces cooperative binding and synergy, supporting a mechanism in which F723-mediated contacts between the P-loop and the allosteric inhibitor are critical for synergy. These structural and mechanistic insights will aid in the identification and development of additional inhibitor combinations with potential clinical value.
- Department of Cancer Biology, Dana-Farber Cancer Institute, Boston, MA, 02215, USA.
Organizational Affiliation: