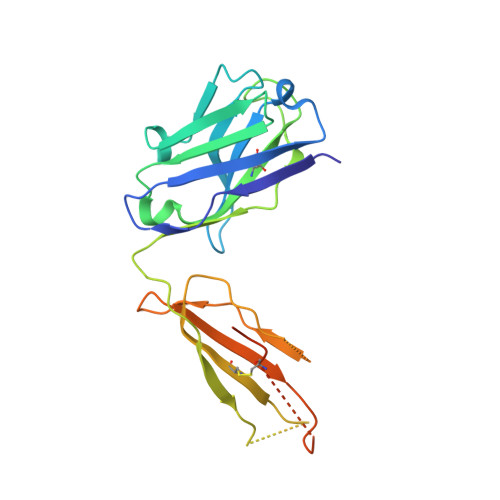
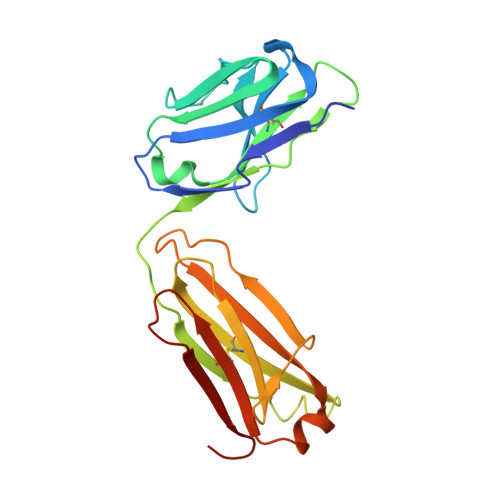
Two human antibodies to a meningococcal serogroup B vaccine antigen enhance binding of complement Factor H by stabilizing the Factor H binding site.
Sands, N.A., Beernink, P.T.(2021) PLoS Pathog 17: e1009655-e1009655
- PubMed: 34125873 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1009655
- Primary Citation Related Structures:
7KE1, 7KET, 7LCV - PubMed Abstract:
Microbial pathogens bind host complement regulatory proteins to evade the immune system. The bacterial pathogen Neisseria meningitidis, or meningococcus, binds several complement regulators, including human Factor H (FH). FH binding protein (FHbp) is a component of two licensed meningococcal vaccines and in mice FHbp elicits antibodies that inhibit binding of FH to FHbp, which defeat the bacterial evasion mechanism. However, humans vaccinated with FHbp develop antibodies that enhance binding of FH to the bacteria, which could limit the effectiveness of the vaccines. In the present study, we show that two vaccine-elicited antibody fragments (Fabs) isolated from different human subjects increase binding of complement FH to meningococcal FHbp by ELISA. The two Fabs have different effects on the kinetics of FH binding to immobilized FHbp as measured by surface plasmon resonance. The 1.7- and 2.0-Å resolution X-ray crystal structures of the Fabs in complexes with FHbp illustrate that the two Fabs bind to similar epitopes on the amino-terminal domain of FHbp, adjacent to the FH binding site. Superposition models of ternary complexes of each Fab with FHbp and FH show that there is likely minimal contact between the Fabs and FH. Collectively, the structures reveal that the Fabs enhance binding of FH to FHbp by altering the conformations and mobilities of two loops adjacent to the FH binding site of FHbp. In addition, the 1.5 Å-resolution structure of one of the isolated Fabs defines the structural rearrangements associated with binding to FHbp. The FH-enhancing human Fabs, which are mirrored in the human polyclonal antibody responses, have important implications for tuning the effectiveness of FHbp-based vaccines.
- Division of Infectious Diseases and Global Health, Department of Pediatrics, School of Medicine, University of California San Francisco, San Francisco, California, United States of America.
Organizational Affiliation: