Structure of a GNAT superfamily PA3944 acetyltransferase in complex with zinc
Czub, M.P., Porebski, P.J., Cymborowski, M., Shabalin, I.G., Minor, W.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Acetyltransferase PA3944 | 194 | Pseudomonas aeruginosa | Mutation(s): 0 Gene Names: PA3944 EC: 2.3.1 | 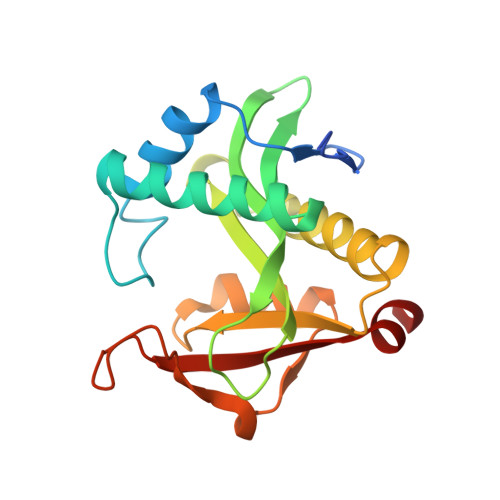 | |
UniProt | |||||
Find proteins for Q9HX72 (Pseudomonas aeruginosa (strain ATCC 15692 / DSM 22644 / CIP 104116 / JCM 14847 / LMG 12228 / 1C / PRS 101 / PAO1)) Explore Q9HX72 Go to UniProtKB: Q9HX72 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q9HX72 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| COA (Subject of Investigation/LOI) Query on COA | E [auth A], J [auth B] | COENZYME A C21 H36 N7 O16 P3 S RGJOEKWQDUBAIZ-IBOSZNHHSA-N |  | ||
| ZN Query on ZN | D [auth A], I [auth B] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| EDO Query on EDO | K [auth B] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| ACT Query on ACT | C [auth A], H [auth B] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| UNX Query on UNX | F [auth A], G [auth A] | UNKNOWN ATOM OR ION X |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 36.423 | α = 98.05 |
| b = 43.801 | β = 109.51 |
| c = 60.793 | γ = 90.1 |
| Software Name | Purpose |
|---|---|
| HKL-3000 | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| HKL-3000 | data reduction |
| MOLREP | phasing |
| Coot | model building |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | -- |