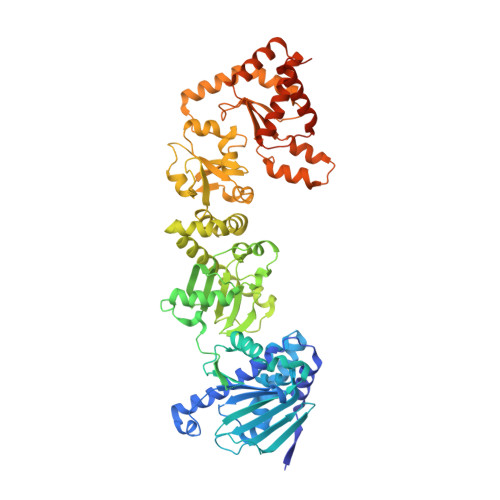
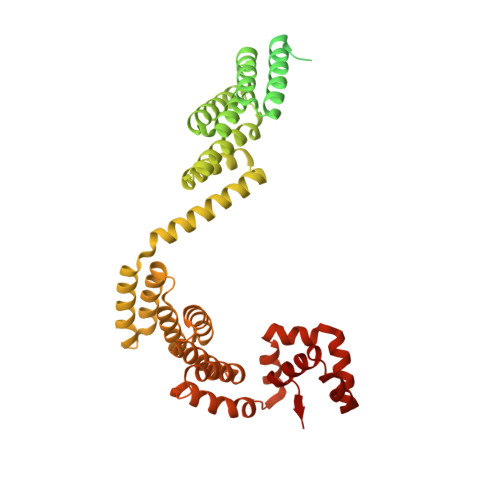
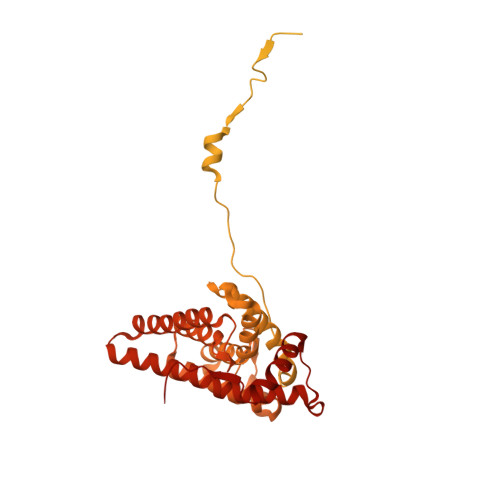
Structure of Hsp90-Hsp70-Hop-GR reveals the Hsp90 client-loading mechanism.
Wang, R.Y., Noddings, C.M., Kirschke, E., Myasnikov, A.G., Johnson, J.L., Agard, D.A.(2022) Nature 601: 460-464
- PubMed: 34937942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-04252-1
- Primary Citation Related Structures:
7KW7 - PubMed Abstract:
Maintaining a healthy proteome is fundamental for the survival of all organisms 1 . Integral to this are Hsp90 and Hsp70, molecular chaperones that together facilitate the folding, remodelling and maturation of the many 'client proteins' of Hsp90 2 . The glucocorticoid receptor (GR) is a model client protein that is strictly dependent on Hsp90 and Hsp70 for activity 3-7 . Chaperoning GR involves a cycle of inactivation by Hsp70; formation of an inactive GR-Hsp90-Hsp70-Hop 'loading' complex; conversion to an active GR-Hsp90-p23 'maturation' complex; and subsequent GR release 8 . However, to our knowledge, a molecular understanding of this intricate chaperone cycle is lacking for any client protein. Here we report the cryo-electron microscopy structure of the GR-loading complex, in which Hsp70 loads GR onto Hsp90, uncovering the molecular basis of direct coordination by Hsp90 and Hsp70. The structure reveals two Hsp70 proteins, one of which delivers GR and the other scaffolds the Hop cochaperone. Hop interacts with all components of the complex, including GR, and poises Hsp90 for subsequent ATP hydrolysis. GR is partially unfolded and recognized through an extended binding pocket composed of Hsp90, Hsp70 and Hop, revealing the mechanism of GR loading and inactivation. Together with the GR-maturation complex structure 9 , we present a complete molecular mechanism of chaperone-dependent client remodelling, and establish general principles of client recognition, inhibition, transfer and activation.
- Department of Biochemistry and Biophysics, University of California San Francisco, San Francisco, CA, USA.
Organizational Affiliation: