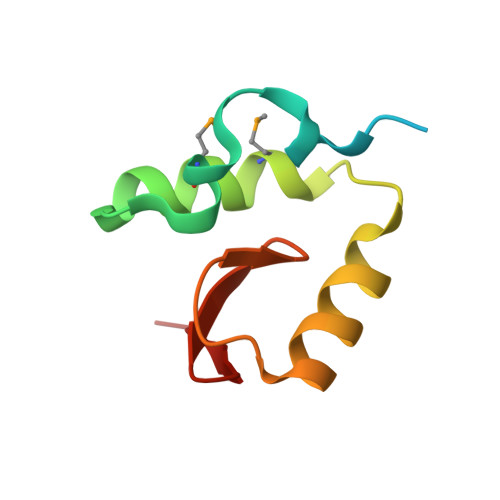
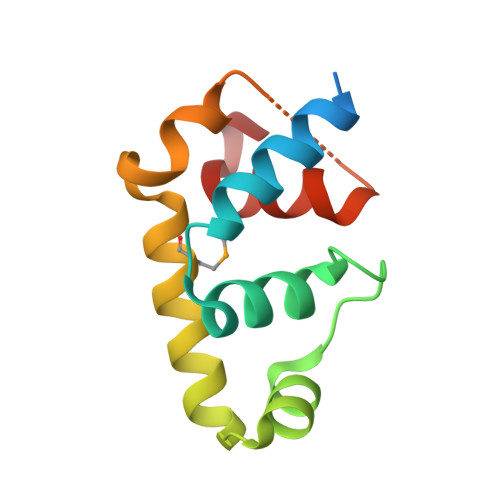
Structural insights into the functional divergence of WhiB-like proteins in Mycobacterium tuberculosis.
Wan, T., Horova, M., Beltran, D.G., Li, S., Wong, H.X., Zhang, L.M.(2021) Mol Cell 81: 2887
- PubMed: 34171298 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2021.06.002
- Primary Citation Related Structures:
7KUF, 7KUG - PubMed Abstract:
WhiB7 represents a distinct subclass of transcription factors in the WhiB-Like (Wbl) family, a unique group of iron-sulfur (4Fe-4S] cluster-containing proteins exclusive to the phylum of Actinobacteria. In Mycobacterium tuberculosis (Mtb), WhiB7 interacts with domain 4 of the primary sigma factor (σ A 4 ) in the RNA polymerase holoenzyme and activates genes involved in multiple drug resistance and redox homeostasis. Here, we report crystal structures of the WhiB7:σ A 4 complex alone and bound to its target promoter DNA at 1.55-Å and 2.6-Å resolution, respectively. These structures show how WhiB7 regulates gene expression by interacting with both σ A 4 and the AT-rich sequence upstream of the -35 promoter DNA via its C-terminal DNA-binding motif, the AT-hook. By combining comparative structural analysis of the two high-resolution σ A 4 -bound Wbl structures with molecular and biochemical approaches, we identify the structural basis of the functional divergence between the two distinct subclasses of Wbl proteins in Mtb.
- Department of Biochemistry, University of Nebraska-Lincoln, Lincoln, NE 68588, USA.
Organizational Affiliation: