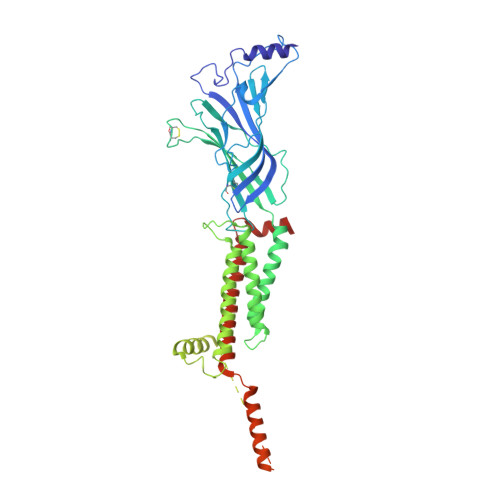
Structure and gating mechanism of the alpha 7 nicotinic acetylcholine receptor.
Noviello, C.M., Gharpure, A., Mukhtasimova, N., Cabuco, R., Baxter, L., Borek, D., Sine, S.M., Hibbs, R.E.(2021) Cell 184: 2121
- PubMed: 33735609 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2021.02.049
- Primary Citation Related Structures:
7KOO, 7KOQ, 7KOX - PubMed Abstract:
The α7 nicotinic acetylcholine receptor plays critical roles in the central nervous system and in the cholinergic inflammatory pathway. This ligand-gated ion channel assembles as a homopentamer, is exceptionally permeable to Ca 2+ , and desensitizes faster than any other Cys-loop receptor. The α7 receptor has served as a prototype for the Cys-loop superfamily yet has proven refractory to structural analysis. We present cryo-EM structures of the human α7 nicotinic receptor in a lipidic environment in resting, activated, and desensitized states, illuminating the principal steps in the gating cycle. The structures also reveal elements that contribute to its function, including a C-terminal latch that is permissive for channel opening, and an anionic ring in the extracellular vestibule that contributes to its high conductance and calcium permeability. Comparisons among the α7 structures provide a foundation for mapping the gating cycle and reveal divergence in gating mechanisms in the Cys-loop receptor superfamily.
- Department of Neuroscience, Peter O'Donnell Jr. Brain Institute, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA; Department of Biophysics, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: