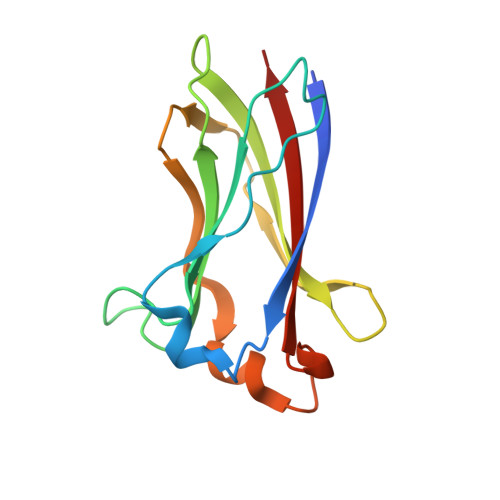
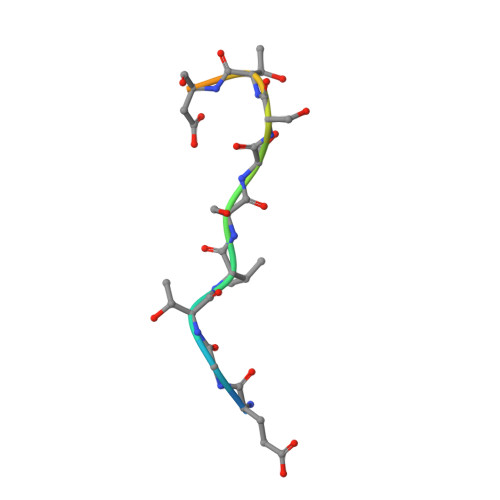
SPOP mutation induces replication over-firing by impairing Geminin ubiquitination and triggers replication catastrophe upon ATR inhibition.
Ma, J., Shi, Q., Cui, G., Sheng, H., Botuyan, M.V., Zhou, Y., Yan, Y., He, Y., Wang, L., Wang, Y., Mer, G., Ye, D., Wang, C., Huang, H.(2021) Nat Commun 12: 5779-5779
- PubMed: 34599168 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-26049-6
- Primary Citation Related Structures:
7KLZ - PubMed Abstract:
Geminin and its binding partner Cdt1 are essential for the regulation of DNA replication. Here we show that the CULLIN3 E3 ubiquitin ligase adaptor protein SPOP binds Geminin at endogenous level and regulates DNA replication. SPOP promotes K27-linked non-degradative poly-ubiquitination of Geminin at lysine residues 100 and 127. This poly-ubiquitination of Geminin prevents DNA replication over-firing by indirectly blocking the association of Cdt1 with the MCM protein complex, an interaction required for DNA unwinding and replication. SPOP is frequently mutated in certain human cancer types and implicated in tumorigenesis. We show that cancer-associated SPOP mutations impair Geminin K27-linked poly-ubiquitination and induce replication origin over-firing and re-replication. The replication stress caused by SPOP mutations triggers replication catastrophe and cell death upon ATR inhibition. Our results reveal a tumor suppressor role of SPOP in preventing DNA replication over-firing and genome instability and suggest that SPOP-mutated tumors may be susceptible to ATR inhibitor therapy.
- Department of Urology, Fudan University Shanghai Cancer Center, Shanghai, 200032, China.
Organizational Affiliation: