Structural Analysis of Toxin-Neutralizing, Single-Domain Antibodies that Bridge Ricin's A-B Subunit Interface.
Rudolph, M.J., Poon, A.Y., Kavaliauskiene, S., Myrann, A.G., Reynolds-Peterson, C., Davis, S.A., Sandvig, K., Vance, D.J., Mantis, N.J.(2021) J Mol Biology 433: 167086-167086
- PubMed: 34089718 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2021.167086
- Primary Citation Related Structures:
7KBI, 7KBK, 7KC9, 7KD0, 7KD2, 7KDM, 7KDU - PubMed Abstract:
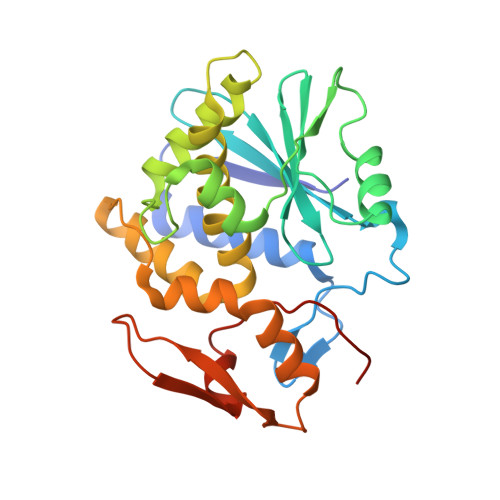
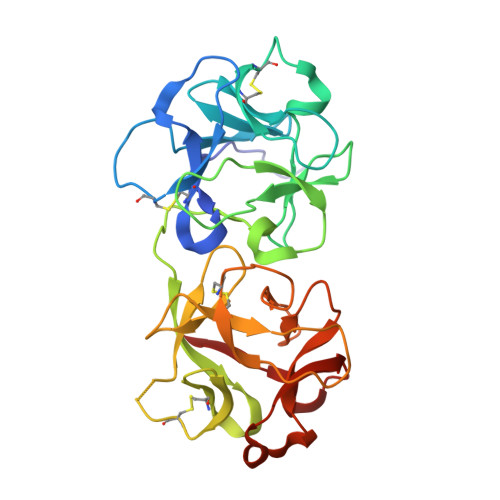
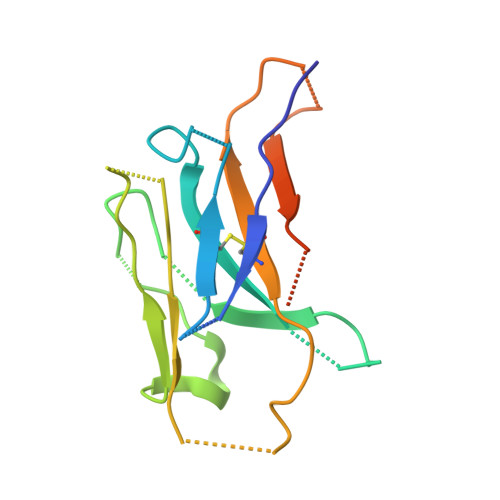
Ricin toxin kills mammalian cells with notorious efficiency. The toxin's B subunit (RTB) is a Gal/GalNAc-specific lectin that attaches to cell surfaces and promotes retrograde transport of ricin's A subunit (RTA) to the trans Golgi network (TGN) and endoplasmic reticulum (ER). RTA is liberated from RTB in the ER and translocated into the cell cytoplasm, where it functions as a ribosome-inactivating protein. While antibodies against ricin's individual subunits have been reported, we now describe seven alpaca-derived, single-domain antibodies (V H Hs) that span the RTA-RTB interface, including four Tier 1 V H Hs with IC 50 values <1 nM. Crystal structures of each V H H bound to native ricin holotoxin revealed three different binding modes, based on contact with RTA's F-G loop (mode 1), RTB's subdomain 2γ (mode 2) or both (mode 3). V H Hs in modes 2 and 3 were highly effective at blocking ricin attachment to HeLa cells and immobilized asialofetuin, due to framework residues (FR3) that occupied the 2γ Gal/GalNAc-binding pocket and mimic ligand. The four Tier 1 V H Hs also interfered with intracellular functions of RTB, as they neutralized ricin in a post-attachment cytotoxicity assay (e.g., the toxin was bound to cell surfaces before antibody addition) and reduced the efficiency of toxin transport to the TGN. We conclude that the RTA-RTB interface is a target of potent toxin-neutralizing antibodies that interfere with both extracellular and intracellular events in ricin's cytotoxic pathway.
- New York Structural Biology Center, New York, NY, USA. Electronic address: mrudolph@nysbc.org.
Organizational Affiliation: