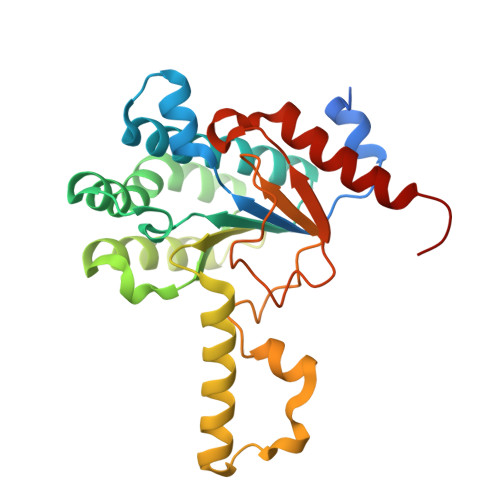
Structural Insights into the Inhibition of Undecaprenyl Pyrophosphate Synthase from Gram-Positive Bacteria.
Workman, S.D., Day, J., Farha, M.A., El Zahed, S.S., Bon, C., Brown, E.D., Organ, M.G., Strynadka, N.C.J.(2021) J Med Chem 64: 13540-13550
- PubMed: 34473495 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.1c00941
- Primary Citation Related Structures:
7JLI, 7JLJ, 7JLM, 7JLR - PubMed Abstract:
The polyprenyl lipid undecaprenyl phosphate (C 55 P) is the universal carrier lipid for the biosynthesis of bacterial cell wall polymers. C 55 P is synthesized in its pyrophosphate form by undecaprenyl pyrophosphate synthase (UppS), an essential cis -prenyltransferase that is an attractive target for antibiotic development. We previously identified a compound (MAC-0547630) that showed promise as a novel class of inhibitor and an ability to potentiate β-lactam antibiotics. Here, we provide a structural model for MAC-0547630's inhibition of UppS and a structural rationale for its enhanced effect on UppS from Bacillus subtilis versus Staphylococcus aureus . We also describe the synthesis of a MAC-0547630 derivative (JPD447), show that it too can potentiate β-lactam antibiotics, and provide a structural rationale for its improved potentiation. Finally, we present an improved structural model of clomiphene's inhibition of UppS. Taken together, our data provide a foundation for structure-guided drug design of more potent UppS inhibitors in the future.
- Department of Biochemistry and Molecular Biology, University of British Columbia, 2350 Health Sciences Mall, Vancouver, British Columbia V6T 1Z3, Canada.
Organizational Affiliation: