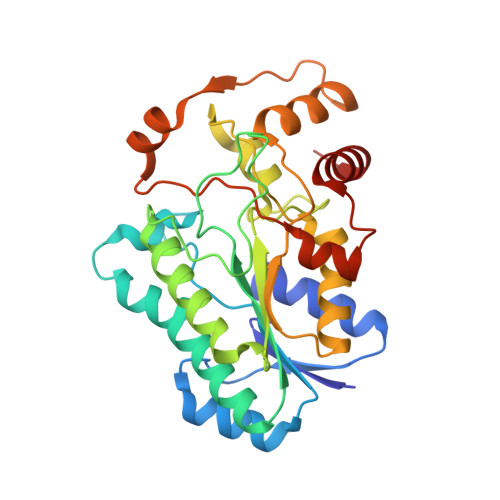
The high-resolution structure of a UDP-L-rhamnose synthase from Acanthamoeba polyphaga Mimivirus.
Bockhaus, N.J., Ferek, J.D., Thoden, J.B., Holden, H.M.(2020) Protein Sci 29: 2164-2174
- PubMed: 32797646 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3928
- Primary Citation Related Structures:
7JID - PubMed Abstract:
For the field of virology, perhaps one of the most paradigm-shifting events so far in the 21st century was the identification of the giant double-stranded DNA virus that infects amoebae. Remarkably, this virus, known as Mimivirus, has a genome that encodes for nearly 1,000 proteins, some of which are involved in the biosynthesis of unusual sugars. Indeed, the virus is coated by a layer of glycosylated fibers that contain d-glucose, N-acetyl-d-glucosamine, l-rhamnose, and 4-amino-4,6-dideoxy-d-glucose. Here we describe a combined structural and enzymological investigation of the protein encoded by the open-reading frame L780, which corresponds to an l-rhamnose synthase. The structure of the L780/NADP + /UDP-l-rhamnose ternary complex was determined to 1.45 Å resolution and refined to an overall R-factor of 19.9%. Each subunit of the dimeric protein adopts a bilobal-shaped appearance with the N-terminal domain harboring the dinucleotide-binding site and the C-terminal domain positioning the UDP-sugar into the active site. The overall molecular architecture of L780 places it into the short-chain dehydrogenase/reductase superfamily. Kinetic analyses indicate that the enzyme can function on either UDP- and dTDP-sugars but displays a higher catalytic efficiency with the UDP-linked substrate. Site-directed mutagenesis experiments suggest that both Cys 108 and Lys 175 play key roles in catalysis. This structure represents the first model of a viral UDP-l-rhamnose synthase and provides new details into these fascinating enzymes.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin, USA.
Organizational Affiliation: