Discovery of a Series of Indane-Containing NBTIs with Activity against Multidrug-Resistant Gram-Negative Pathogens.
Cumming, J.G., Kreis, L., Kuhne, H., Wermuth, R., Vercruysse, M., Kramer, C., Rudolph, M.G., Xu, Z.(2023) ACS Med Chem Lett 14: 993-998
- PubMed: 37465290 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.3c00187
- Primary Citation Related Structures:
7FVS, 7FVT - PubMed Abstract:
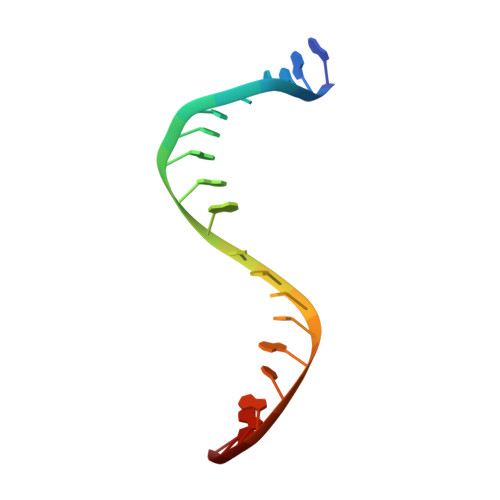
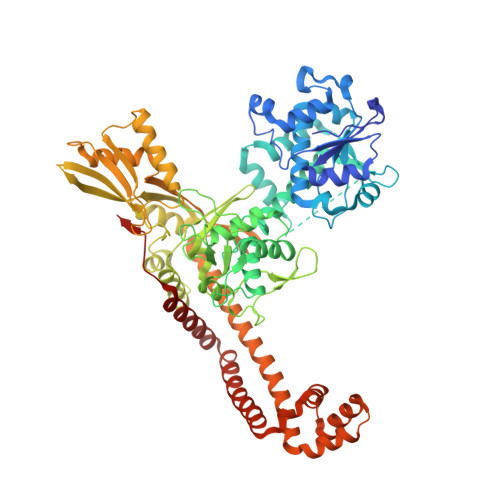
The rise of multidrug-resistant (MDR) Gram-negative bacteria is a major global health problem necessitating the discovery of new classes of antibiotics. Novel bacterial topoisomerase inhibitors (NBTIs) target the clinically validated bacterial type II topoisomerases with a distinct binding site and mechanism of action to fluoroquinolone antibiotics, thus avoiding cross-resistance to this drug class. Here we report the discovery of a series of NBTIs incorporating a novel indane DNA binding moiety. X-ray cocrystal structures of compounds 2 and 17a bound to Staphylococcus aureus DNA gyrase-DNA were determined, revealing specific interactions with the enzyme binding pocket at the GyrA dimer interface and a long-range electrostatic interaction between the basic amine in the linker and the carboxylate of Asp83. Exploration of the structure-activity relationship within the series led to the identification of lead compound 18c , which showed potent broad-spectrum activity against a panel of MDR Gram-negative bacteria.
- Roche Pharma Research & Early Development, F. Hoffmann-La Roche Ltd., CH-4070 Basel, Switzerland.
Organizational Affiliation: