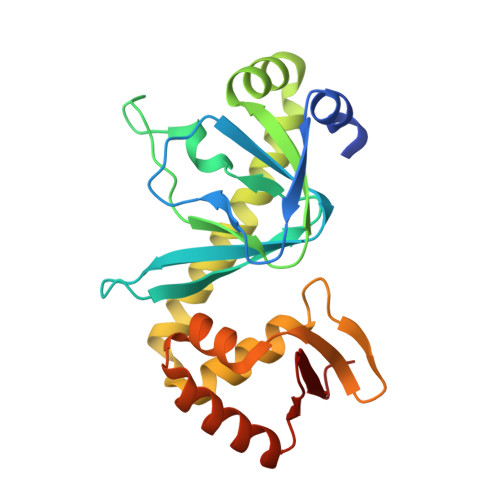
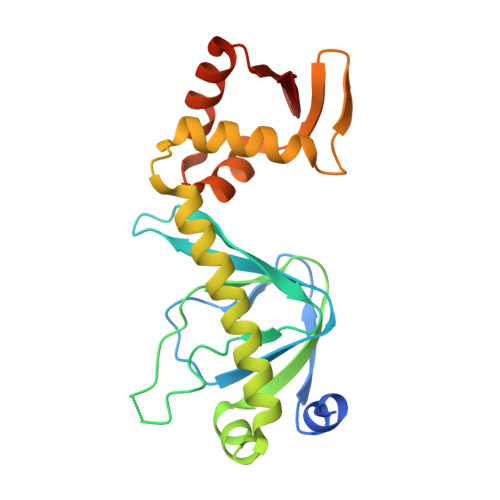
Structural basis for the inhibitory mechanism of auranofin and gold(I) analogues against Pseudomonas aeruginosa global virulence factor regulator Vfr.
Zhang, Y., Chew, B.L.A., Wang, J., Yuan, M., Yam, J.K.H., Luo, D., Yang, L.(2023) Comput Struct Biotechnol J 21: 2137-2146
- PubMed: 37007650 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.csbj.2023.03.013
- Primary Citation Related Structures:
7FEW, 7FF0, 7FF8, 7FF9 - PubMed Abstract:
Pseudomonas aeruginosa is a leading cause of hospital-acquired infections. Treatment of P. aeruginosa infections is difficult given its multiple virulence mechanisms, intrinsic antibiotic resistance mechanisms, and biofilm-forming ability. Auranofin, an approved oral gold compound for rheumatoid arthritis treatment, was recently reported to inhibit the growth of multiple bacterial species. Here, we identify P. aeruginosa 's global virulence factor regulator Vfr as one target of auranofin. We report the mechanistic insights into the inhibitory mechanism of auranofin and gold(I) analogues to Vfr through structural, biophysical, and phenotypic inhibition studies. This work suggests that auranofin and gold(I) analogues have potential to be developed as anti-virulence drugs against P. aeruginosa.
- Shenzhen Third People's Hospital, The Second Affiliated Hospital of Southern University of Science and Technology, National Clinical Research Center for Infectious Disease, Shenzhen 518112, China.
Organizational Affiliation: