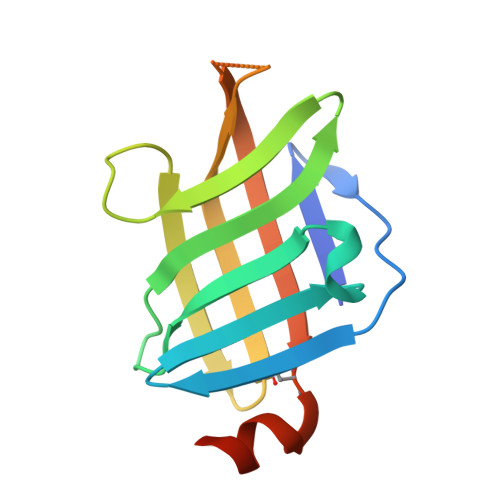
High-resolution crystal structure of LpqH, an immunomodulatory surface lipoprotein of Mycobacterium tuberculosis reveals a distinct fold and a conserved cleft on its surface.
Chatterjee, S., Kundapura, S.V., Basak, A.J., Mukherjee, D., Dash, S., Ganguli, N., Das, A.K., Mukherjee, G., Samanta, D., Ramagopal, U.A.(2022) Int J Biol Macromol 210: 494-503