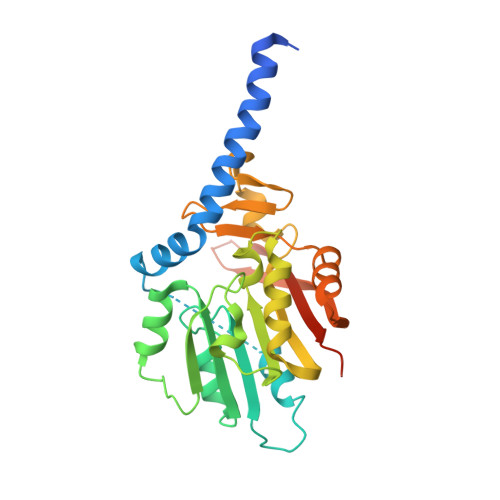
Structural basis for METTL6-mediated m3C RNA methylation.
Li, S., Zhou, H., Liao, S., Wang, X., Zhu, Z., Zhang, J., Xu, C.(2021) Biochem Biophys Res Commun 589: 159-164
- PubMed: 34922197
- DOI: https://doi.org/10.1016/j.bbrc.2021.12.013
- Primary Citation of Related Structures:
7F1E - PubMed Abstract:
RNA modifications play important roles in mediating the biological functions of RNAs. 3-methylcytidine (m3C), albeit less abundant, is found to exist extensively in tRNAs, rRNAs and mRNAs. Human METTL6 is a m 3 C methyltransferase for tRNAs, including tRNA SER(UGA) . We solved the structure of human METTL6 in the presence of S-adenosyl-L-methionine and found by enzyme assay that recombinant human METTL6 is active towards tRNA SER(UGA) . Structural analysis indicated the detailed interactions between S-adenosyl-L-methionine and METTL6, and suggested potential tRNA binding region on the surface of METTL6. The structural research, complemented by biochemistry enzyme assay, will definitely shed light on the design of potent inhibitors for METTL6 in near future.
- MOE Key Laboratory for Membraneless Organelles & Cellular Dynamics, Hefei National Laboratory for Physical Sciences at the Microscale, School of Life Sciences, Division of Life Sciences and Medicine, University of Science and Technology of China, 230027, Hefei, PR China.
Organizational Affiliation: