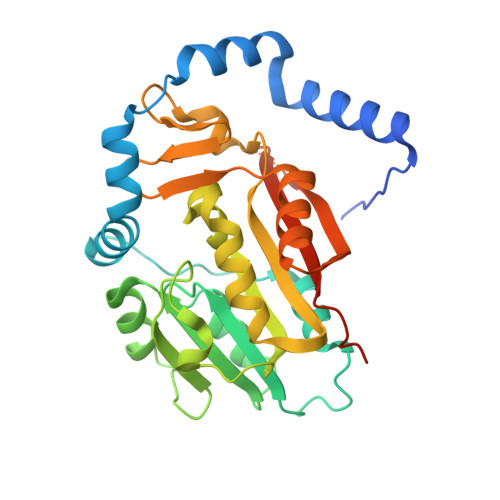
Crystal structure of human METTL6, the m 3 C methyltransferase.
Chen, R., Zhou, J., Liu, L., Mao, X.L., Zhou, X., Xie, W.(2021) Commun Biol 4: 1361-1361
- PubMed: 34862464
- DOI: https://doi.org/10.1038/s42003-021-02890-9
- Primary Citation Related Structures:
7EZG - PubMed Abstract:
In tRNA, the epigenetic m 3 C modification at position 32 in the anticodon loop is highly conserved in eukaryotes, which maintains the folding and basepairing functions of the anticodon. However, the responsible enzymes METTL2 and METTL6 were identified only in recent years. The loss of human METTL6 (hMETTL6) affects the translational process and proteostasis in cells, while in mESCs cells, it leads to defective pluripotency potential. Despite its important functions, the catalytic mechanism of the C32 methylation by this enzyme is poorly understood. Here we present the 1.9 Å high-resolution crystal structure of hMETTL6 bound by SAH. The key residues interacting with the ligand were identified and their roles were confirmed by ITC. We generated a docking model for the hMETTL6-SAH-CMP ternary complex. Interestingly, the CMP molecule binds into a cavity in a positive patch with the base ring pointing to the inside, suggesting a flipped-base mechanism for methylation. We further generated a model for the quaternary complex with tRNA Ser as a component, which reasonably explained the biochemical behaviors of hMETTL6. Taken together, our crystallographic and biochemical studies provide important insight into the molecular recognition mechanism by METTL6 and may aid in the METTL-based rational drug design in the future.
- MOE Key Laboratory of Gene Function and Regulation, State Key Laboratory for Biocontrol, School of Life Sciences, The Sun Yat-Sen University, Guangzhou, Guangdong, 510006, People's Republic of China.
Organizational Affiliation: