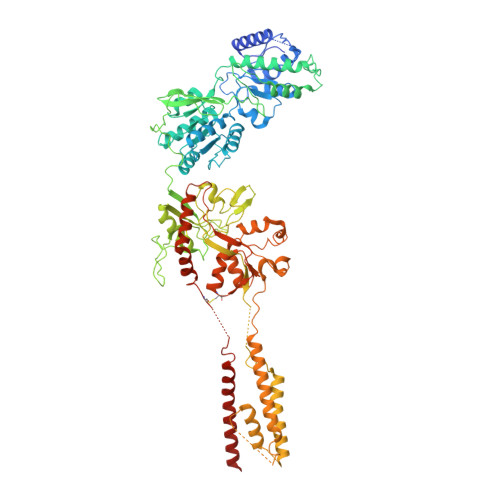
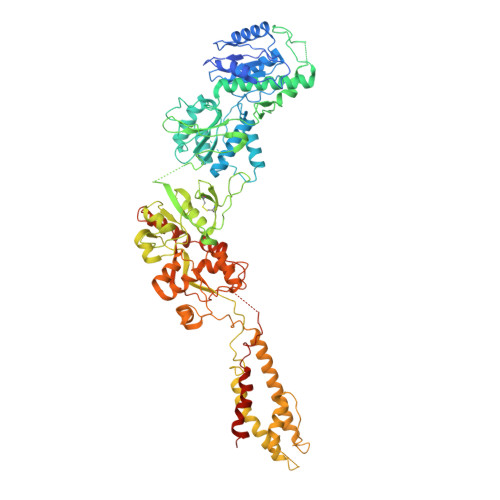
Structural basis of ketamine action on human NMDA receptors.
Zhang, Y., Ye, F., Zhang, T., Lv, S., Zhou, L., Du, D., Lin, H., Guo, F., Luo, C., Zhu, S.(2021) Nature 596: 301-305
- PubMed: 34321660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-021-03769-9
- Primary Citation Related Structures:
7EU7, 7EU8 - PubMed Abstract:
Ketamine is a non-competitive channel blocker of N-methyl-D-aspartate (NMDA) receptors 1 . A single sub-anaesthetic dose of ketamine produces rapid (within hours) and long-lasting antidepressant effects in patients who are resistant to other antidepressants 2,3 . Ketamine is a racemic mixture of S- and R-ketamine enantiomers, with S-ketamine isomer being the more active antidepressant 4 . Here we describe the cryo-electron microscope structures of human GluN1-GluN2A and GluN1-GluN2B NMDA receptors in complex with S-ketamine, glycine and glutamate. Both electron density maps uncovered the binding pocket for S-ketamine in the central vestibule between the channel gate and selectivity filter. Molecular dynamics simulation showed that S-ketamine moves between two distinct locations within the binding pocket. Two amino acids-leucine 642 on GluN2A (homologous to leucine 643 on GluN2B) and asparagine 616 on GluN1-were identified as key residues that form hydrophobic and hydrogen-bond interactions with ketamine, and mutations at these residues reduced the potency of ketamine in blocking NMDA receptor channel activity. These findings show structurally how ketamine binds to and acts on human NMDA receptors, and pave the way for the future development of ketamine-based antidepressants.
- Institute of Neuroscience, State Key Laboratory of Neuroscience, CAS Center for Excellence in Brain Science and Intelligence Technology, Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: