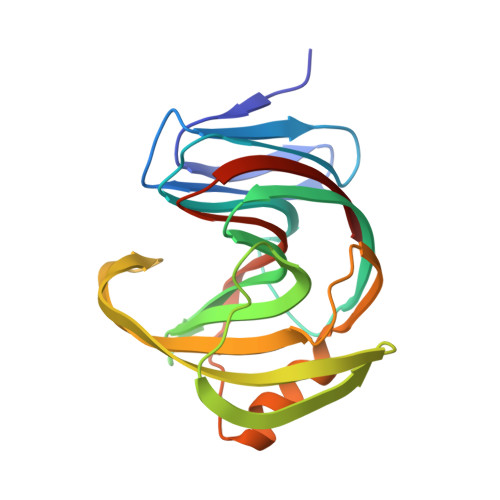
Characterization and structural analysis of a thermophilic GH11 xylanase from compost metatranscriptome.
Yi, Y., Xu, S., Kovalevsky, A., Zhang, X., Liu, D., Wan, Q.(2021) Appl Microbiol Biotechnol 105: 7757-7767
- PubMed: 34553251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s00253-021-11587-2
- Primary Citation Related Structures:
7EO6 - PubMed Abstract:
Xylanase is efficient for xylan degradation and widely applied in industries. We found a GH11 family xylanase (Xyn11A) with high thermostability and catalytic activity from compost metatranscriptome. This xylanase has the optimal reaction temperature at 80 °C with the activity of 2907.3 U/mg. The X-ray crystallographic structure shows a typical "right hand" architecture, which is the characteristics of the GH11 family enzymes. Comparing it with the mesophilic XYN II, a well-studied GH11 xylanase from Trichoderma reesei, Xyn11A is more compact with more H-bonds. Our mutagenic results show that the electrostatic interactions in the thumb and palm region of Xyn11A could result in its high thermostability and activity. Introducing a disulfide bond at the N-terminus further increased its optimal reaction temperature to 90 °C with augmented activity. KEY POINTS: • A hyperthermophilic xylanase with high activity was discovered using the metatranscriptomic method. • The mechanisms of thermophilicity and high activity were revealed using X-ray crystallography, mutagenesis, and molecular dynamics simulations. • The thermostability and activity were further improved by introducing a disulfide bond.
- College of Science, Nanjing Agricultural University, Nanjing, 210095, People's Republic of China.
Organizational Affiliation: