Identification and structural analysis of a thermophilic beta-1,3-glucanase from compost
Feng, J., Xu, S., Feng, R., Kovalevsky, A., Zhang, X., Liu, D., Wan, Q.(2021) Bioresour Bioprocess 8: 102-102
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 1,3-beta-glucanase | 283 | Actinomycetes bacterium | Mutation(s): 0 Gene Names: DIU60_04100 | 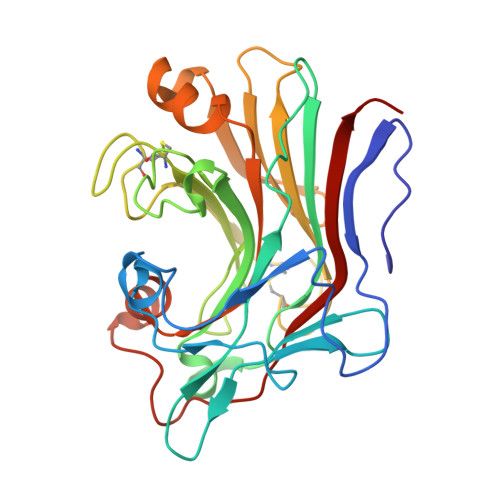 | |
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| TRS Download:Ideal Coordinates CCD File | B [auth A] | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL C4 H12 N O3 LENZDBCJOHFCAS-UHFFFAOYSA-O |  | ||
| MG Download:Ideal Coordinates CCD File | C [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 60.397 | α = 90 |
| b = 61.01 | β = 90 |
| c = 70.672 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| HKL-2000 | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-3000 | data reduction |
| HKL-3000 | data collection |