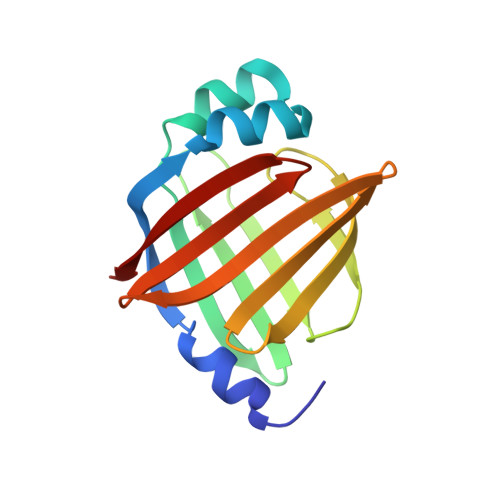
Crystal structure of human brain-type fatty acid-binding protein FABP7 complexed with palmitic acid.
Nam, K.H.(2021) Acta Crystallogr D Struct Biol 77: 954-965
- PubMed: 34196621 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798321005763
- Primary Citation Related Structures:
7E25 - PubMed Abstract:
The brain-type fatty acid-binding protein FABP7, which is expressed in astrocytes and neural progenitors, is a member of the intracellular lipid-binding protein family. This protein is not only involved in various cellular functions such as metabolism, inflammation and energy homeostasis, but also in diseases such as cognitive disorders and tumors. Structures of unsaturated fatty acids, such as oleic acid (OA) and docosahexaenoic acid (DHA), bound to FABP7 have been elucidated; however, structures of saturated fatty acids bound to FABP7 remain unknown. To better understand fatty acid recognition, here the crystal structure of human brain-type fatty acid-binding protein FABP7 complexed with palmitic acid (PA), a saturated fatty acid, is reported at a resolution of 1.6 Å. The PA bound to the fatty acid-binding pocket of FABP7 assumed a U-shaped conformation. The carboxylate moiety of PA interacted with Tyr129, Arg127 and, via a water bridge, with Arg107 and Thr54, whereas its aliphatic chain was stabilized by hydrophobic interactions with Met21, Leu24, Thr30, Thr37, Pro39, Phe58 and Asp77. Structural comparison showed that PA, OA and DHA exhibited unique binding conformations in the fatty acid-binding pocket, stabilized by distinct amino-acid interactions. The binding of PA to FABP7 exhibits a unique binding conformation when compared with other human FABPs (FABP3-FABP5 and FABP8) expressed in other tissues. Based on the crystal and fatty acid structures, it was suggested that PA, which prefers a linear form in nature, required a greater conformational change in its aliphatic chain to bind to the fatty acid-binding pocket in a U-shaped conformation, compared with the cis configurations of OA or DHA. This, together with the length of the aliphatic chain, was considered to be one of the factors determining the binding affinity of PA to FABP7. These results provide a better understanding of fatty acid recognition by FABP7 and expand the knowledge of the binding of PA to FABPs.
- Department of Life Science, Pohang University of Science and Technology, Pohang, Republic of Korea.
Organizational Affiliation: