2-Cyanoisonicotinamide Conjugation: A Facile Approach to Generate Potent Peptide Inhibitors of the Zika Virus Protease.
Patil, N.A., Quek, J.P., Schroeder, B., Morewood, R., Rademann, J., Luo, D., Nitsche, C.(2021) ACS Med Chem Lett 12: 732-737
- PubMed: 34055219 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.0c00657
- Primary Citation Related Structures:
7DOC - PubMed Abstract:
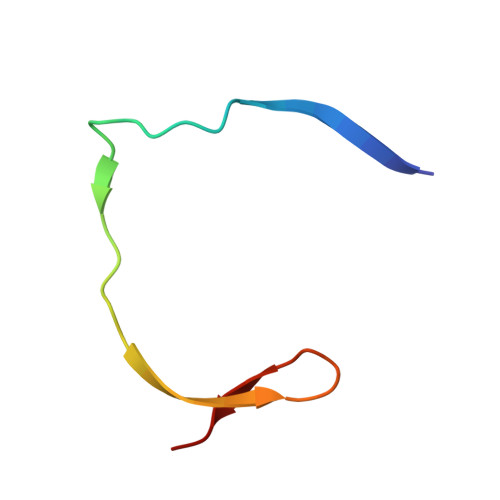
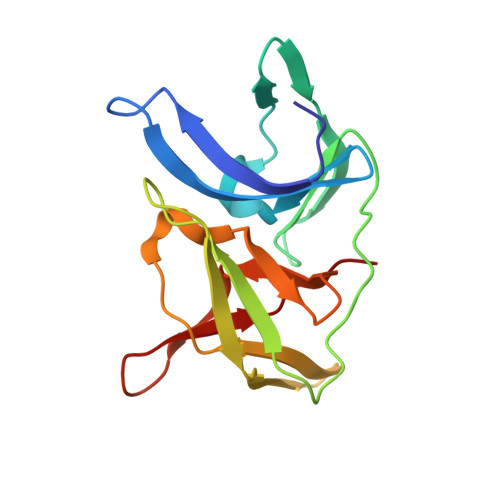
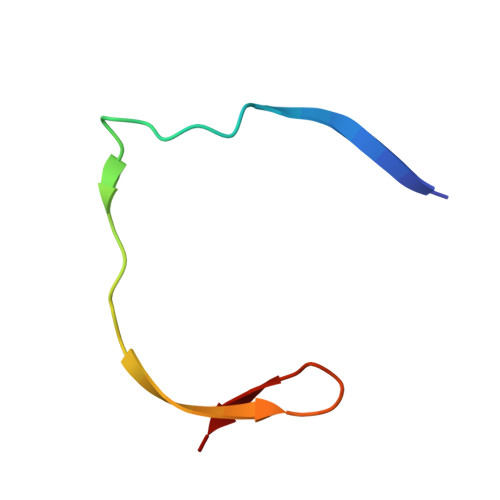
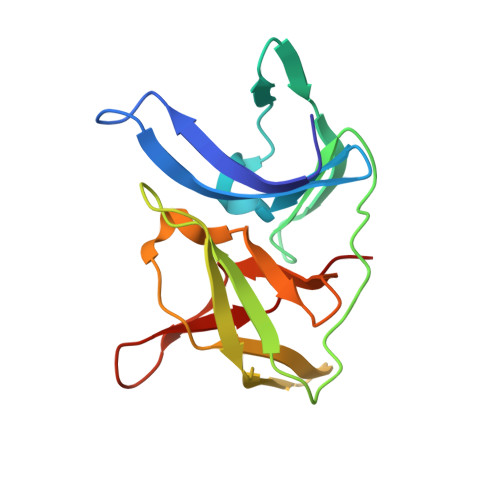
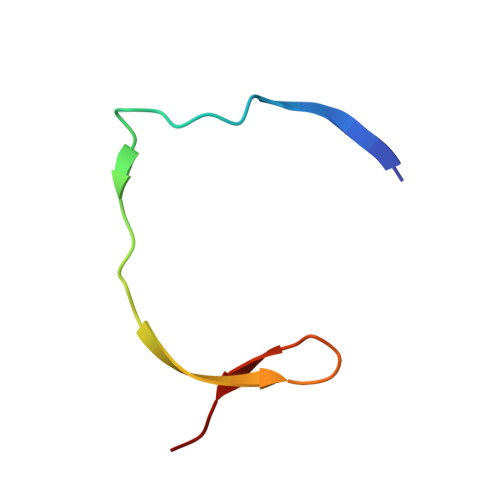
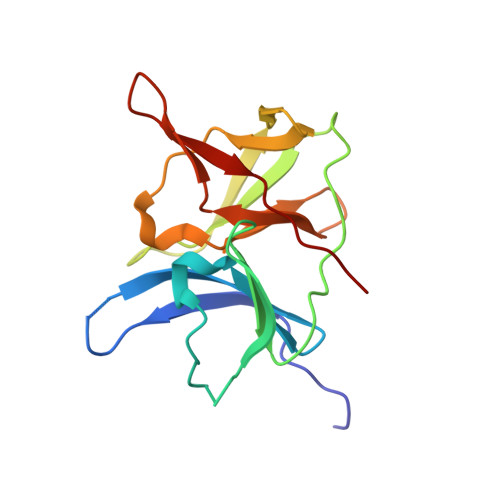
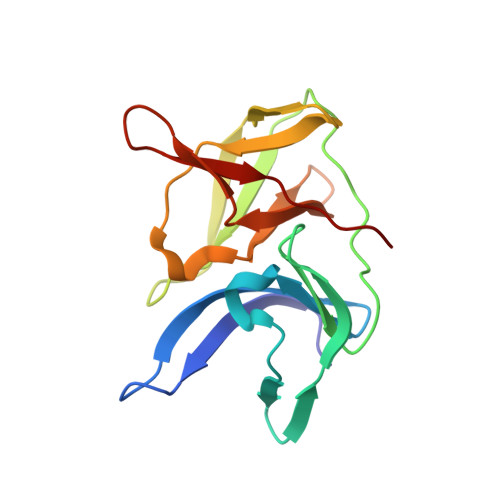
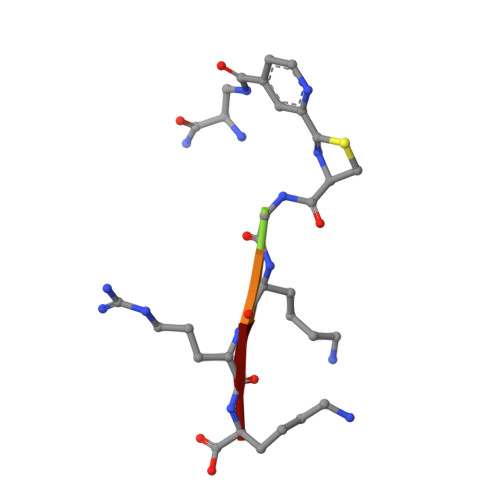
The rapid generation and modification of macrocyclic peptides in medicinal chemistry is an ever-growing area that can present various synthetic challenges. The reaction between N-terminal cysteine and 2-cyanoisonicotinamide is a new biocompatible click reaction that allows rapid access to macrocyclic peptides. Importantly, 2-cyanoisonicotinamide can be attached to different linkers directly during solid-phase peptide synthesis. The synthesis involves only commercially available precursors, allowing for a fully automated process. We demonstrate the approach for four cyclic peptide ligands of the Zika virus protease NS2B-NS3. Although all peptides display the substrate recognition motif, the activity strongly depends on the linker length, with the shortest cyclization linker corresponding to highest activity ( K i = 0.64 μM). The most active cyclic peptide displays affinity 78 times higher than that of its linear analogue. We solved a crystal structure of the proteolytically cleaved ligand and synthesized it by applying the presented chemistry to peptide ligation.
- Biomedicine Discovery Institute, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: