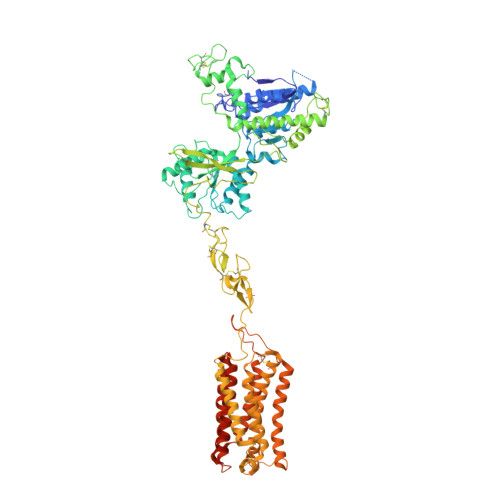
Structural insights into the activation initiation of full-length mGlu1.
Zhang, J., Qu, L., Wu, L., Tang, X., Luo, F., Xu, W., Xu, Y., Liu, Z.J., Hua, T.(2021) Protein Cell 12: 662-667
- PubMed: 33278019 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-020-00808-5
- Primary Citation Related Structures:
7DGD, 7DGE - iHuman Institute, ShanghaiTech University, Shanghai, 201210, China.
Organizational Affiliation: