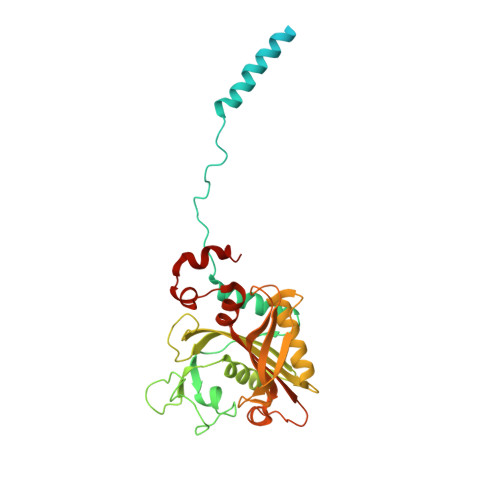
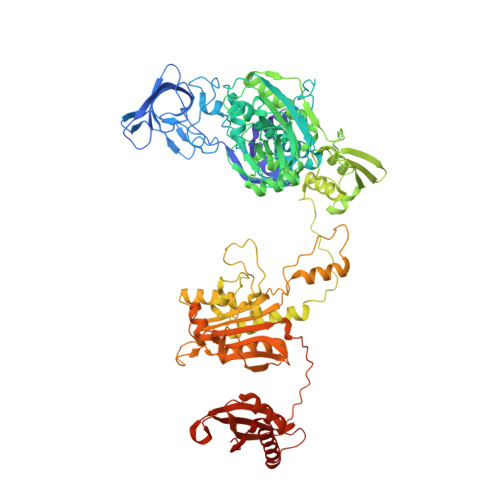
Re-discovery of PF-3845 as a new chemical scaffold inhibiting phenylalanyl-tRNA synthetase in Mycobacterium tuberculosis .
Wang, H., Xu, M., Engelhart, C.A., Zhang, X., Yan, B., Pan, M., Xu, Y., Fan, S., Liu, R., Xu, L., Hua, L., Schnappinger, D., Chen, S.(2021) J Biological Chem
- PubMed: 33397709 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.RA120.016477
- Primary Citation Related Structures:
7DAW, 7DB7, 7DB8 - PubMed Abstract:
Mycobacteria tuberculosis (Mtb) remains the deadliest pathogenic bacteria worldwide. The search for new antibiotics to treat drug-sensitive as well as drug-resistant tuberculosis has become a priority. The essential enzyme phenylalanyl-tRNA synthetase (PheRS) is an antibacterial drug target because of the large differences between bacterial and human PheRS counterparts. In a high-throughput screening of 2148 bioactive compounds, PF-3845, which is a known inhibitor of human fatty acid amide hydrolase (FAAH), was identified inhibiting Mtb PheRS at K i ~0.73 ± 0.06 µM. The inhibition mechanism was studied with enzyme kinetics, protein structural modelling and crystallography, in comparison to a PheRS inhibitor of the noted phenyl-thiazolylurea-sulfonamide class. The 2.3-Å crystal structure of Mtb PheRS in complex with PF-3845 revealed its novel binding mode, in which a trifluoromethyl-pyridinylphenyl group occupies the Phe pocket while a piperidine-piperazine urea group binds into the ATP pocket through an interaction network enforced by a sulfate ion. It represents the first non-nucleoside bi-substrate competitive inhibitor of bacterial PheRS. PF-3845 inhibits the in vitro growth of Mtb H37Rv at ~24 µM, and the potency of PF-3845 increased against Mtb pheS-FDAS, suggesting on target activity in mycobacterial whole cells. PF-3845 does not inhibit human cytoplasmic or mitochondrial PheRS in biochemical assay, which can be explained from the crystal structures. Further medicinal chemistry efforts focused on the piperidine-piperazine urea moiety may result in the identification of a selective antibacterial lead compound.
- Global Health Drug Discovery Institute, China.
Organizational Affiliation: