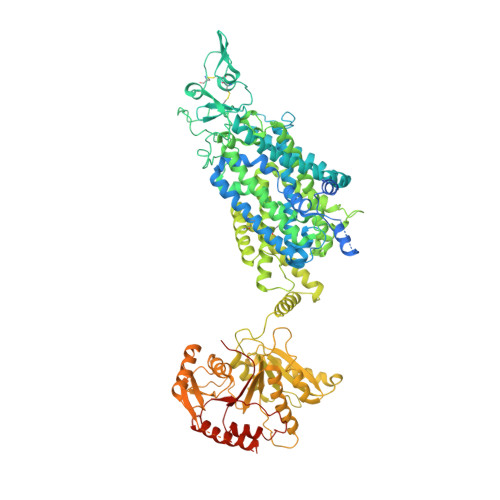
Structures and an activation mechanism of human potassium-chloride cotransporters.
Xie, Y., Chang, S., Zhao, C., Wang, F., Liu, S., Wang, J., Delpire, E., Ye, S., Guo, J.(2020) Sci Adv 6
- PubMed: 33310850 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abc5883
- Primary Citation Related Structures:
7D8Z, 7D90, 7D99 - PubMed Abstract:
Potassium-chloride cotransporters KCC1 to KCC4 mediate the coupled export of potassium and chloride across the plasma membrane and play important roles in cell volume regulation, auditory system function, and γ-aminobutyric acid (GABA) and glycine-mediated inhibitory neurotransmission. Here, we present 2.9- to 3.6-Å resolution structures of full-length human KCC2, KCC3, and KCC4. All three KCCs adopt a similar overall architecture, a domain-swap dimeric assembly, and an inward-facing conformation. The structural and functional studies reveal that one unexpected N-terminal peptide binds at the cytosolic facing cavity and locks KCC2 and KCC4 at an autoinhibition state. The C-terminal domain (CTD) directly interacts with the N-terminal inhibitory peptide, and the relative motions between the CTD and the transmembrane domain (TMD) suggest that CTD regulates KCCs' activities by adjusting the autoinhibitory effect. These structures provide the first glimpse of full-length structures of KCCs and an autoinhibition mechanism among the amino acid-polyamine-organocation transporter superfamily.
- Department of Biophysics, and Department of Pathology of Sir Run Run Shaw Hospital, Zhejiang University School of Medicine, Hangzhou 310058, China.
Organizational Affiliation: