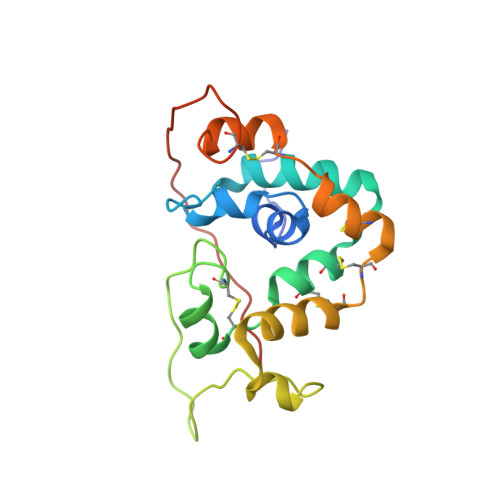
Solution structure of Gaussia Luciferase with five disulfide bonds and identification of a putative coelenterazine binding cavity by heteronuclear NMR.
Wu, N., Kobayashi, N., Tsuda, K., Unzai, S., Saotome, T., Kuroda, Y., Yamazaki, T.(2020) Sci Rep 10: 20069-20069
- PubMed: 33208800 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-76486-4
- Primary Citation Related Structures:
7D2O - PubMed Abstract:
Gaussia luciferase (GLuc) is a small luciferase (18.2 kDa; 168 residues) and is thus attracting much attention as a reporter protein, but the lack of structural information is hampering further application. Here, we report the first solution structure of a fully active, recombinant GLuc determined by heteronuclear multidimensional NMR. We obtained a natively folded GLuc by bacterial expression and efficient refolding using a Solubility Enhancement Petide (SEP) tag. Almost perfect assignments of GLuc's 1 H, 13 C and 15 N backbone signals were obtained. GLuc structure was determined using CYANA, which automatically identified over 2500 NOEs of which > 570 were long-range. GLuc is an all-alpha-helix protein made of nine helices. The region spanning residues 10-18, 36-81, 96-145 and containing eight out of the nine helices was determined with a C α -atom RMSD of 1.39 Å ± 0.39 Å. The structure of GLuc is novel and unique. Two homologous sequential repeats form two anti-parallel bundles made by 4 helices and tied together by three disulfide bonds. The N-terminal helix 1 is grabbed by these 4 helices. Further, we found a hydrophobic cavity where several residues responsible for bioluminescence were identified in previous mutational studies, and we thus hypothesize that this is a catalytic cavity, where the hydrophobic coelenterazine binds and the bioluminescence reaction takes place.
- College of Food and Bioengineering, Zhengzhou University of Light Industry, 136 Kexue Road, Zhengzhou, 450001, People's Republic of China.
Organizational Affiliation: