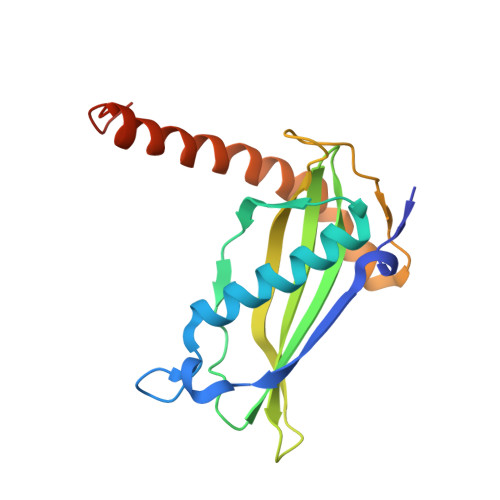
Structural basis for nucleotide-independent regulation of acyl-CoA thioesterase from Bacillus cereus ATCC 14579.
Park, J., Kim, Y.J., Lee, D., Kim, K.J.(2020) Int J Biol Macromol 170: 390-396
- PubMed: 33383082
- DOI: https://doi.org/10.1016/j.ijbiomac.2020.12.174
- Primary Citation of Related Structures:
7CZ3 - PubMed Abstract:
Acyl-CoA thioesterase is an enzyme that catalyzes the cleavage of thioester bonds and regulates the cellular concentrations of CoASH, fatty acids, and acyl-CoA. In this study, we report the crystal structure of acyl-CoA thioesterase from Bacillus cereus ATCC 14579 (BcACT1) complexed with the CoA product. BcACT1 possesses a monomeric structure of a hotdog-fold and forms a hexamer via the trimerization of three dimers. We identified the active site of BcACT1 and revealed that residues Asn23 and Asp38 are crucial for enzyme catalysis, indicating that BcACT1 belongs to the TE6 family. We also propose that BcACT1 might undergo an open-closed conformational change on the acyl-CoA binding pocket upon binding of the acyl-CoA substrate. Interestingly, the BcACT1 variants with dramatically increased activities were obtained during the site-directed mutagenesis experiments to confirm the residues involved in CoA binding. Finally, we found that BcACT1 is not nucleotide-regulated and suggest that the length and shape of the additional α2-helix are crucial in determining a regulation mode by nucleotides.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daegu 41566, Republic of Korea; KNU Institute for Microorganisms, Kyungpook National University, Daegu 41566, Republic of Korea.
Organizational Affiliation: