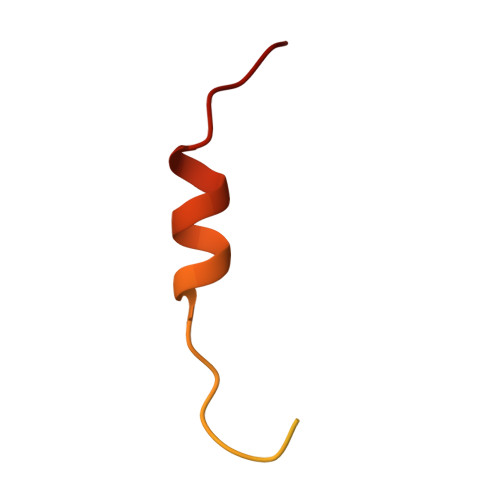
Mechanism of karyopherin-beta 2 binding and nuclear import of ALS variants FUS(P525L) and FUS(R495X).
Gonzalez, A., Mannen, T., Cagatay, T., Fujiwara, A., Matsumura, H., Niesman, A.B., Brautigam, C.A., Chook, Y.M., Yoshizawa, T.(2021) Sci Rep 11: 3754-3754
- PubMed: 33580145 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-021-83196-y
- Primary Citation Related Structures:
7CYL - PubMed Abstract:
Mutations in the RNA-binding protein FUS cause familial amyotropic lateral sclerosis (ALS). Several mutations that affect the proline-tyrosine nuclear localization signal (PY-NLS) of FUS cause severe juvenile ALS. FUS also undergoes liquid-liquid phase separation (LLPS) to accumulate in stress granules when cells are stressed. In unstressed cells, wild type FUS resides predominantly in the nucleus as it is imported by the importin Karyopherin-β2 (Kapβ2), which binds with high affinity to the C-terminal PY-NLS of FUS. Here, we analyze the interactions between two ALS-related variants FUS(P525L) and FUS(R495X) with importins, especially Kapβ2, since they are still partially localized to the nucleus despite their defective/missing PY-NLSs. The crystal structure of the Kapβ2·FUS(P525L) PY-NLS complex shows the mutant peptide making fewer contacts at the mutation site, explaining decreased affinity for Kapβ2. Biochemical analysis revealed that the truncated FUS(R495X) protein, although missing the PY-NLS, can still bind Kapβ2 and suppresses LLPS. FUS(R495X) uses its C-terminal tandem arginine-glycine-glycine regions, RGG2 and RGG3, to bind the PY-NLS binding site of Kapβ2 for nuclear localization in cells when arginine methylation is inhibited. These findings suggest the importance of the C-terminal RGG regions in nuclear import and LLPS regulation of ALS variants of FUS that carry defective PY-NLSs.
- Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, TX, USA.
Organizational Affiliation: