The ligand-free structure of human PPARgamma LBD
Jang, D.M., Han, B.W.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Peroxisome proliferator-activated receptor gamma | 283 | Homo sapiens | Mutation(s): 1 Gene Names: PPARG, NR1C3 | 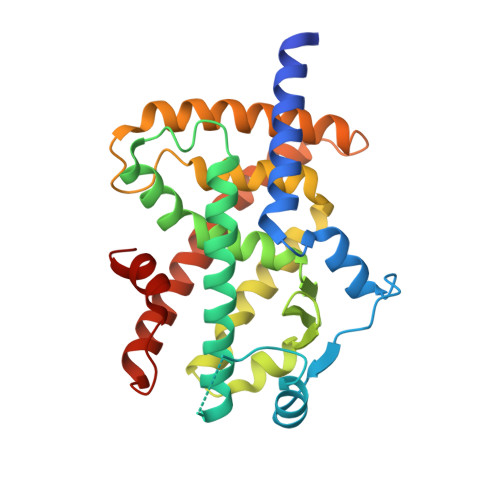 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P37231 GTEx: ENSG00000132170 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P37231 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 16-mer peptide from Nuclear receptor coactivator 1 | 16 | Homo sapiens | Mutation(s): 0 EC: 2.3.1.48 | 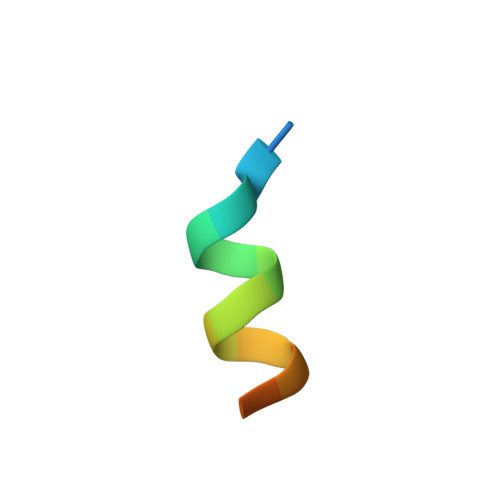 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q15788 GTEx: ENSG00000084676 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15788 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| MLI (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | C [auth A] | MALONATE ION C3 H2 O4 OFOBLEOULBTSOW-UHFFFAOYSA-L |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 130.396 | α = 90 |
| b = 51.997 | β = 90 |
| c = 54.052 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| MOLREP | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Research Foundation (NRF, Korea) | Korea, Republic Of | 2011-0030001 |