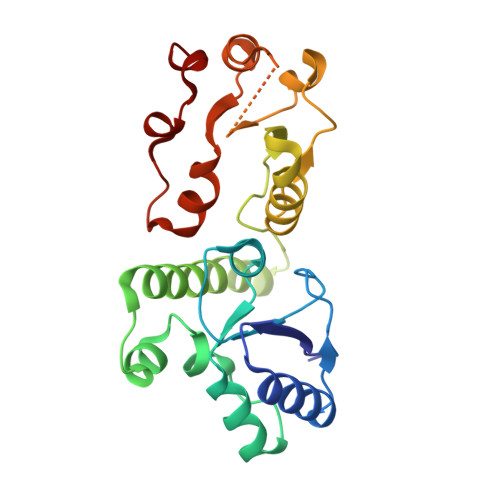
PHF8-promoted TOPBP1 demethylation drives ATR activation and preserves genome stability.
Ma, S., Cao, C., Che, S., Wang, Y., Su, D., Liu, S., Gong, W., Liu, L., Sun, J., Zhao, J., Wang, Q., Song, N., Ge, T., Guo, Q., Tian, S., Chen, C.D., Zhang, T., Wang, J., Ding, X., Yang, F., Ying, G., Yang, J., Zhang, K., Zhu, Y., Yao, Z., Yang, N., Shi, L.(2021) Sci Adv 7
- PubMed: 33952527 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abf7684
- Primary Citation Related Structures:
7CMZ - PubMed Abstract:
The checkpoint kinase ATR [ATM (ataxia-telangiectasia mutated) and rad3-related] is a master regulator of DNA damage response. Yet, how ATR activity is regulated remains to be investigated. We report here that histone demethylase PHF8 (plant homeodomain finger protein 8) plays a key role in ATR activation and replication stress response. Mechanistically, PHF8 interacts with and demethylates TOPBP1 (DNA topoisomerase 2-binding protein 1), an essential allosteric activator of ATR, under unperturbed conditions, but replication stress results in PHF8 phosphorylation and dissociation from TOPBP1. Consequently, hypomethylated TOPBP1 facilitates RAD9 (RADiation sensitive 9) binding and chromatin loading of the TOPBP1-RAD9 complex to fully activate ATR and thus safeguard the genome and protect cells against replication stress. Our study uncovers a demethylation and phosphorylation code that controls the assembly of TOPBP1-scaffolded protein complex, and provides molecular insight into non-histone methylation switch in ATR activation.
- State Key Laboratory of Experimental Hematology, 2011 Collaborative Innovation Center of Tianjin for Medical Epigenetics, Key Laboratory of Breast Cancer Prevention and Therapy (Ministry of Education), Key Laboratory of Immune Microenvironment and Disease (Ministry of Education), Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Tianjin Medical University Cancer Institute and Hospital, Tianjin Medical University General Hospital, Tianjin Medical University, Tianjin 300070, China.
Organizational Affiliation: