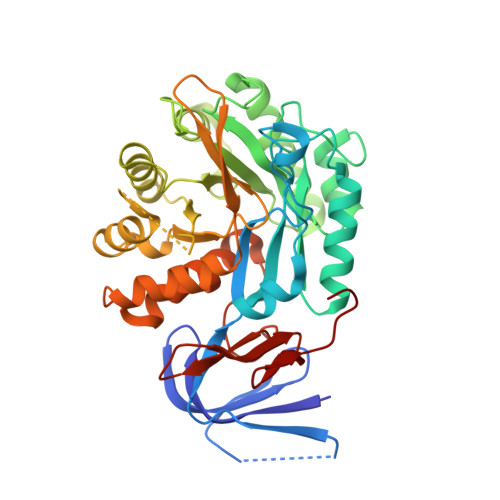
Functional Characterization of Primordial Protein Repair Enzyme M38 Metallo-Peptidase From Fervidobacterium islandicum AW-1.
La, J.W., Dhanasingh, I., Jang, H., Lee, S.H., Lee, D.W.(2020) Front Mol Biosci 7: 600634-600634
- PubMed: 33392259 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmolb.2020.600634
- Primary Citation Related Structures:
7CDH, 7CF6 - PubMed Abstract:
The NA23_RS08100 gene of Fervidobacterium islandicum AW-1 encodes a keratin-degrading β-aspartyl peptidase ( Fi BAP) that is highly expressed under starvation conditions. Herein, we expressed the gene in Escherichia coli , purified the recombinant enzyme to homogeneity, and investigated its function. The 318 kDa recombinant Fi BAP enzyme exhibited maximal activity at 80°C and pH 7.0 in the presence of Zn 2+ . Size-exclusion chromatography revealed that the native enzyme is an octamer comprising a tetramer of dimers; this was further supported by determination of its crystal structure at 2.6 Å resolution. Consistently, the structure of Fi BAP revealed three additional salt bridges in each dimer, involving 12 ionic interactions that might contribute to its high thermostability. In addition, the co-crystal structure containing the substrate analog N -carbobenzoxy-β-Asp-Leu at 2.7 Å resolution revealed binuclear Zn 2+ -mediated substrate binding, suggesting that Fi BAP is a hyperthermophilic type-I IadA, in accordance with sequence-based phylogenetic analysis. Indeed, complementation of a Leu auxotrophic E. coli mutant strain (Δ iadA and Δ leuB ) with Fi BAP enabled the mutant strain to grow on isoAsp-Leu peptides. Remarkably, LC-MS/MS analysis of soluble keratin hydrolysates revealed that Fi BAP not only cleaves the C-terminus of isoAsp residues but also has a relatively broad substrate specificity toward α-peptide bonds. Moreover, heat shock-induced protein aggregates retarded bacterial growth, but expression of BAP alleviated the growth defect by degrading damaged proteins. Taken together, these results suggest that the viability of hyperthermophiles under stressful conditions may rely on the activity of BAP within cellular protein repair systems.
- Department of Biotechnology, Yonsei University, Seoul, South Korea.
Organizational Affiliation: