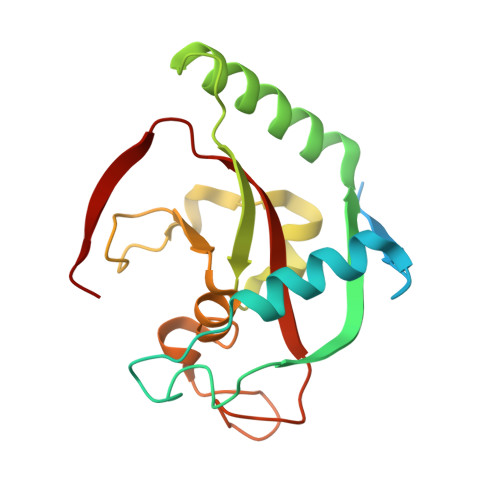
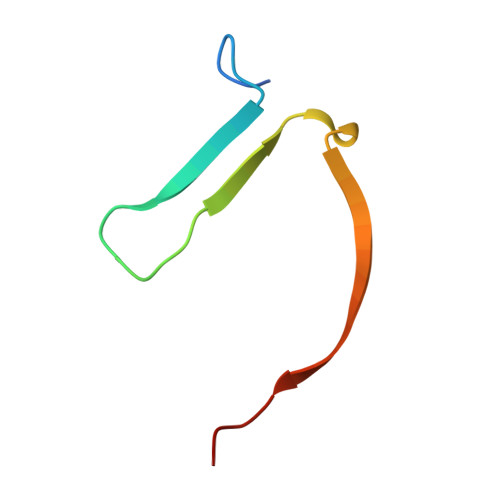
The dual pocket binding novel tankyrase inhibitor K-476 enhances the efficacy of immune checkpoint inhibitor by attracting CD8 + T cells to tumors.
Kinosada, H., Okada-Iwasaki, R., Kunieda, K., Suzuki-Imaizumi, M., Takahashi, Y., Miyagi, H., Suzuki, M., Motosawa, K., Watanabe, M., Mie, M., Ishii, T., Ishida, H., Saito, J.I., Nakai, R.(2021) Am J Cancer Res 11: 264-276
- PubMed: 33520373 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
7CE4 - PubMed Abstract:
The Wnt/β-catenin pathway, which is associated with disease progression, is activated in many cancers. Tankyrase (TNKS) has received attention as a target molecule for Wnt/β-catenin pathway inhibition. We identified K-476, a novel TNKS inhibitor, a dual pocket binder that binds to both the nicotinamide and ADP-ribose pockets. In a human colon cancer cell line, K-476 specifically and potently inhibited TNKS and led to stabilization of the Axin protein, resulting in Wnt/β-catenin pathway suppression. Aberrant Wnt/β-catenin pathway activation was recently reported as a possible mechanism of ineffectiveness in immune checkpoint inhibitor (ICI) treatment. Because the Wnt/β-catenin pathway activation causes dendritic cell inactivation and suppresses chemokine production, resulting in a paucity of CD8 + T cells in tumor tissue, which is an important effector of ICIs. Thus, TNKS inhibitors may enhance the efficacy of ICIs. To examine whether K-476 enhances the antitumor effect of anti-PD-L1 antibodies, K-476 was administered orally with an anti-PD-L1 antibody to melanoma-bearing C57BL/6J mice. Although K-476 was ineffective as a monotherapy, it significantly enhanced the antitumor effect in combination with anti-PD-L1 antibody. In mice, intra-tumor infiltration of CD8 + T cells was increased by combination treatment. K-476 upregulated the chemokine expression (e.g., Ccl3 and Ccl4 ), which attracted CD8 + T cells. This was considered to contribute to the increased CD8 + T cells in the tumor microenvironment. Furthermore, while the potential gastrointestinal toxicity of TNKS inhibitors has been reported, it was not observed at effective doses. Thus, K-476 could be an attractive therapeutic option to enhance the efficacy of ICIs.
- R&D Division, Kyowa Kirin Co., Ltd. Japan.
Organizational Affiliation: