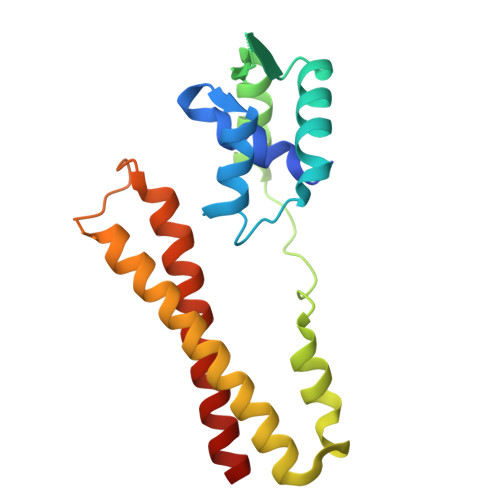
Apo structure of the transcriptional regulator PadR from Bacillus subtilis: Structural dynamics and conserved Y70 residue.
Park, S.C., Song, W.S., Yoon, S.I.(2020) Biochem Biophys Res Commun 530: 215-221
- PubMed: 32828288 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.06.135
- Primary Citation Related Structures:
7CBV - PubMed Abstract:
PadR is a bacterial transcriptional regulator that controls the expression of phenolic acid decarboxylase (PadC) in response to phenolic acids to prevent their toxic effects. During transcriptional repression, PadR associates with the operator sequence at the promoter site of the padC gene. However, when phenolic acids are present, PadR directly binds the phenolic acids and undergoes an interdomain rearrangement to dissociate from the operator DNA. To further examine the structural dynamics of PadR, we determined the apo structure of Bacillus subtilis PadR. Apo-PadR exhibits significant interdomain flexibility and adopts structures that are similar to the phenolic acid-bound PadR structures but distinct from the DNA-bound structure, suggesting that apo-PadR can bind phenolic acids without substantial structural rearrangement. Furthermore, we identified the Y70 residue of PadR as the most conserved residue in the PadR family. PadR Y70 displays similar conformations irrespective of the associated partners, and its conformation is conserved in diverse PadR family members. The Y70 residue is surrounded by the key DNA-binding entities of PadR and is required to optimally arrange them for operator DNA recognition by PadR. PadR Y70 also plays a critical role in protein stability based on the results of a denaturation assay. These observations suggest that PadR Y70 is a canonical residue of the PadR family that contributes to protein stability and DNA binding.
- Division of Biomedical Convergence, College of Biomedical Science, Kangwon National University, Chuncheon, 24341, Republic of Korea.
Organizational Affiliation: