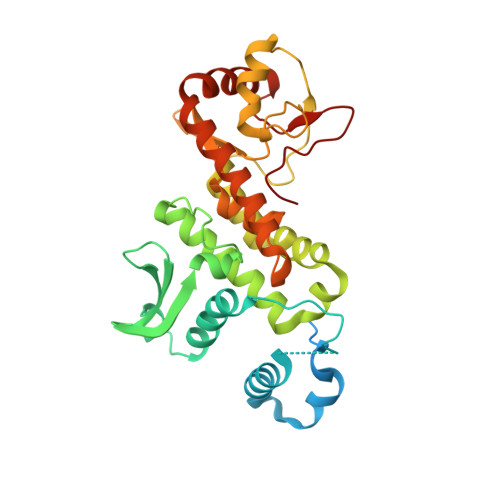
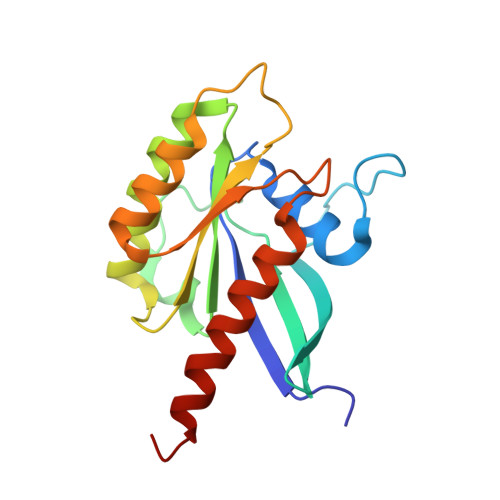
The Salmonella effector protein SopD targets Rab8 to positively and negatively modulate the inflammatory response.
Lian, H., Jiang, K., Tong, M., Chen, Z., Liu, X., Galan, J.E., Gao, X.(2021) Nat Microbiol 6: 658-671
- PubMed: 33603205 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-021-00866-3
- Primary Citation Related Structures:
7BWT - PubMed Abstract:
The food-borne bacterial pathogen Salmonella Typhimurium uses a type III protein secretion system to deliver multiple proteins into host cells. These secreted effectors modulate the functions of host cells and activate specific signalling cascades that result in the production of pro-inflammatory cytokines and intestinal inflammation. Some of the Salmonella-encoded effectors counteract this inflammatory response and help to preserve host homeostasis. Here, we demonstrate that the Salmonella effector protein SopD, which is required for pathogenesis, functions to both activate and inhibit the inflammatory response by targeting the Rab8 GTPase, which is a negative regulator of inflammation. We show that SopD has GTPase-activating protein activity for Rab8 and, therefore, inhibits this GTPase and stimulates inflammation. We also show that SopD activates Rab8 by displacing it from its cognate guanosine dissociation inhibitor, resulting in the stimulation of a signalling cascade that suppresses inflammation. We solved the crystal structure of SopD in association with Rab8 to a resolution of 2.3 Å, which reveals a unique contact interface that underlies these complex interactions. These findings show the remarkable evolution of a bacterial effector protein to exert both agonistic and antagonistic activities towards the same host cellular target to modulate the inflammatory response.
- Department of Microbial Pathogenesis, Yale University School of Medicine, New Haven, CT, USA.
Organizational Affiliation: