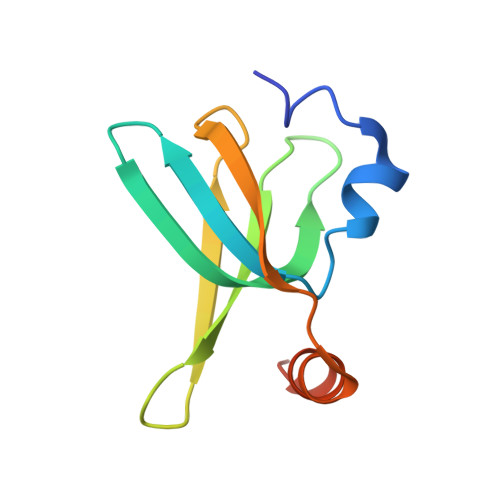
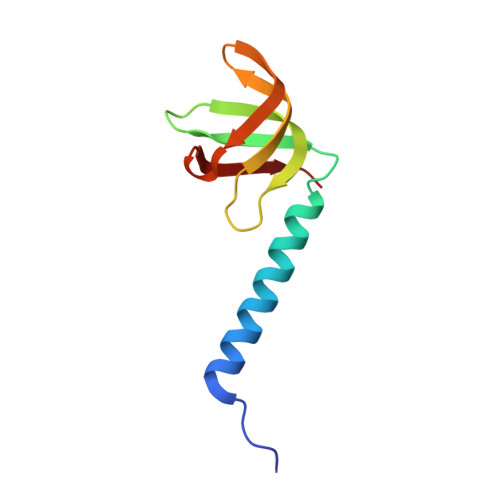
Identification and structural analysis of the Schizosaccharomyces pombe SMN complex.
Veepaschit, J., Viswanathan, A., Bordonne, R., Grimm, C., Fischer, U.(2021) Nucleic Acids Res 49: 7207-7223
- PubMed: 33754639 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab158
- Primary Citation Related Structures:
7BB3, 7BBL - PubMed Abstract:
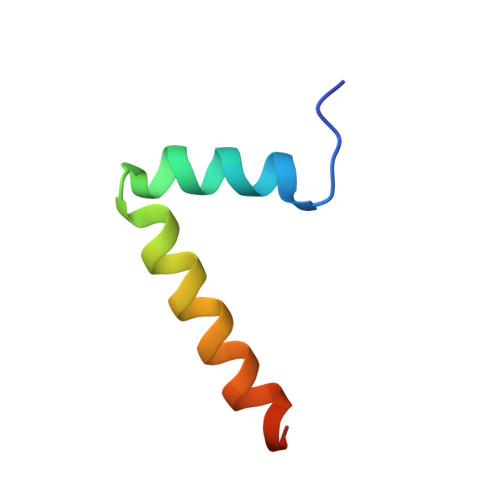
The macromolecular SMN complex facilitates the formation of Sm-class ribonucleoproteins involved in mRNA processing (UsnRNPs). While biochemical studies have revealed key activities of the SMN complex, its structural investigation is lagging behind. Here we report on the identification and structural determination of the SMN complex from the lower eukaryote Schizosaccharomyces pombe, consisting of SMN, Gemin2, 6, 7, 8 and Sm proteins. The core of the SMN complex is formed by several copies of SMN tethered through its C-terminal alpha-helices arranged with alternating polarity. This creates a central platform onto which Gemin8 binds and recruits Gemins 6 and 7. The N-terminal parts of the SMN molecules extrude via flexible linkers from the core and enable binding of Gemin2 and Sm proteins. Our data identify the SMN complex as a multivalent hub where Sm proteins are collected in its periphery to allow their joining with UsnRNA.
- Department of Biochemistry, Biocenter, University of Würzburg, Würzburg 97074, Germany.
Organizational Affiliation: