Crystal structure of MurE from E.coli
Koekemoer, L., Steindel, M., Fairhead, M., Talon, R., Douangamath, A., Arrowsmith, C.H., Edwards, A.M., Bountra, C., von Delft, F., Krojer, T.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| UDP-N-acetylmuramoyl-L-alanyl-D-glutamate-2,6-diaminopimelate ligase | 496 | Escherichia coli K-12 | Mutation(s): 0 Gene Names: murE, b0085, JW0083 EC: 6.3.2.13 | 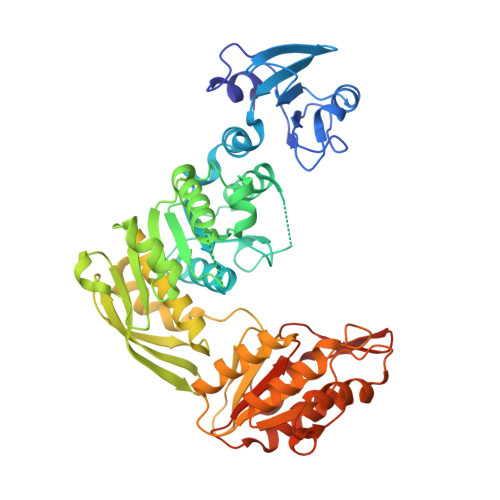 | |
UniProt | |||||
Find proteins for P22188 (Escherichia coli (strain K12)) Explore P22188 Go to UniProtKB: P22188 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P22188 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| JHP (Subject of Investigation/LOI) Query on JHP | C [auth A], D [auth B] | 4-chloro-N-cyclopentyl-1-methyl-1H-pyrazole-3-carboxamide C10 H14 Cl N3 O AEWMHZKRUKXXFH-UHFFFAOYSA-N |  | ||
| IPA Query on IPA | E [auth B] | ISOPROPYL ALCOHOL C3 H8 O KFZMGEQAYNKOFK-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 58.517 | α = 96.93 |
| b = 58.546 | β = 91.45 |
| c = 74.194 | γ = 104.74 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| xia2 | data reduction |
| xia2 | data scaling |
| PHASER | phasing |