Uncovering an allosteric mode of action for a selective inhibitor of human Bloom syndrome protein.
Chen, X., Ali, Y.I., Fisher, C.E., Arribas-Bosacoma, R., Rajasekaran, M.B., Williams, G., Walker, S., Booth, J.R., Hudson, J.J., Roe, S.M., Pearl, L.H., Ward, S.E., Pearl, F.M., Oliver, A.W.(2021) Elife 10
- PubMed: 33647232 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.65339
- Primary Citation Related Structures:
7AUC, 7AUD - PubMed Abstract:
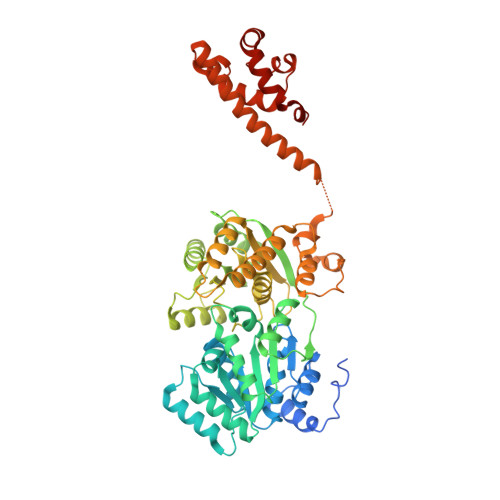
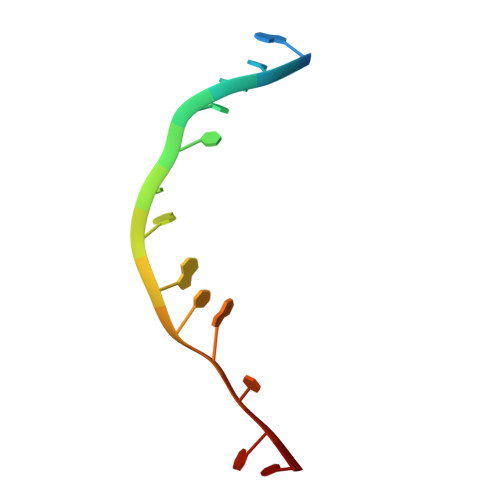
BLM (Bloom syndrome protein) is a RECQ-family helicase involved in the dissolution of complex DNA structures and repair intermediates. Synthetic lethality analysis implicates BLM as a promising target in a range of cancers with defects in the DNA damage response; however, selective small molecule inhibitors of defined mechanism are currently lacking. Here, we identify and characterise a specific inhibitor of BLM's ATPase-coupled DNA helicase activity, by allosteric trapping of a DNA-bound translocation intermediate. Crystallographic structures of BLM-DNA-ADP-inhibitor complexes identify a hitherto unknown interdomain interface, whose opening and closing are integral to translocation of ssDNA, and which provides a highly selective pocket for drug discovery. Comparison with structures of other RECQ helicases provides a model for branch migration of Holliday junctions by BLM.
- Cancer Research UK DNA Repair Enzymes Group, Genome Damage and Stability Centre, School of Life Sciences, University of Sussex, Falmer, United Kingdom.
Organizational Affiliation: