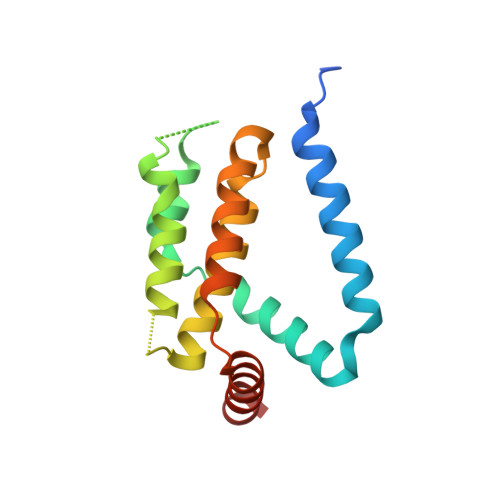
Crystal structures of ORFV125 provide insight into orf virus-mediated inhibition of apoptosis.
Suraweera, C.D., Hinds, M.G., Kvansakul, M.(2020) Biochem J 477: 4527-4541
- PubMed: 33175095 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20200776
- Primary Citation Related Structures:
7ADS, 7ADT - PubMed Abstract:
Premature apoptosis of cells is a strategy utilized by multicellular organisms to counter microbial threats. Orf virus (ORFV) is a large double-stranded DNA virus belonging to the poxviridae. ORFV encodes for an apoptosis inhibitory protein ORFV125 homologous to B-cell lymphoma 2 or Bcl-2 family proteins, which has been shown to inhibit host cell encoded pro-apoptotic Bcl-2 proteins. However, the structural basis of apoptosis inhibition by ORFV125 remains to be clarified. We show that ORFV125 is able to bind to a range of peptides spanning the BH3 motif of human pro-apoptotic Bcl-2 proteins including Bax, Bak, Puma and Hrk with modest to weak affinity. We then determined the crystal structures of ORFV125 alone as well as bound to the highest affinity ligand Bax BH3 motif. ORFV125 adopts a globular Bcl-2 fold comprising 7 α-helices, and utilizes the canonical Bcl-2 binding groove to engage pro-apoptotic host cell Bcl-2 proteins. In contrast with a previously predicted structure, ORFV125 adopts a domain-swapped dimeric topology, where the α1 helix from one protomer is swapped into a neighbouring unit. Furthermore, ORFV125 differs from the conserved architecture of the Bcl-2 binding groove and instead of α3 helix forming one of the binding groove walls, ORFV125 utilizes an extended α2 helix that comprises the equivalent region of helix α3. This results in a subtle variation of previously observed dimeric Bcl-2 architectures in other poxvirus and human encoded Bcl-2 proteins. Overall, our results provide a structural and mechanistic basis for orf virus-mediated inhibition of host cell apoptosis.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, Victoria 3086, Australia.
Organizational Affiliation: