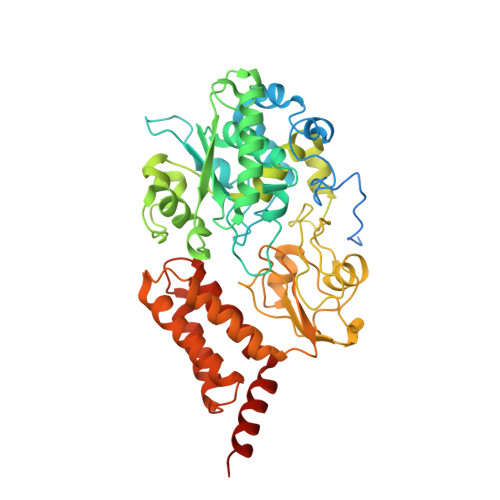
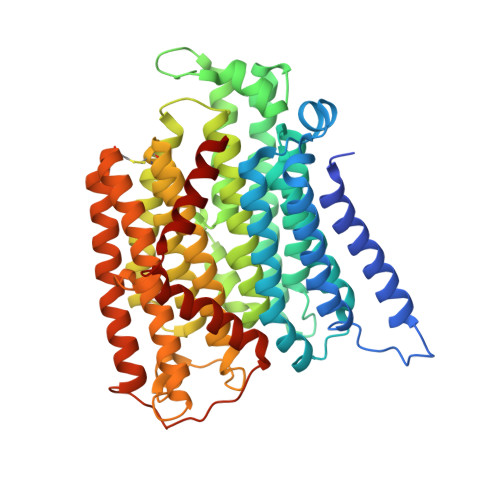
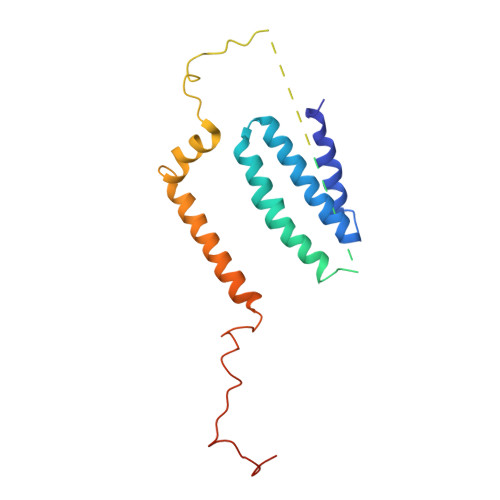
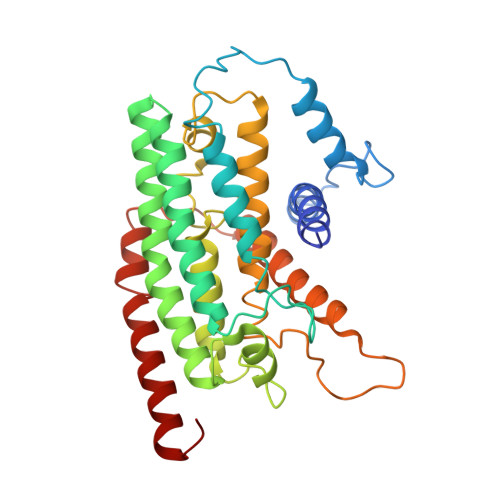
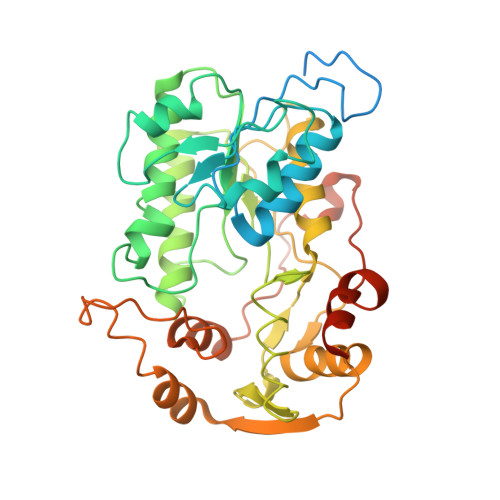
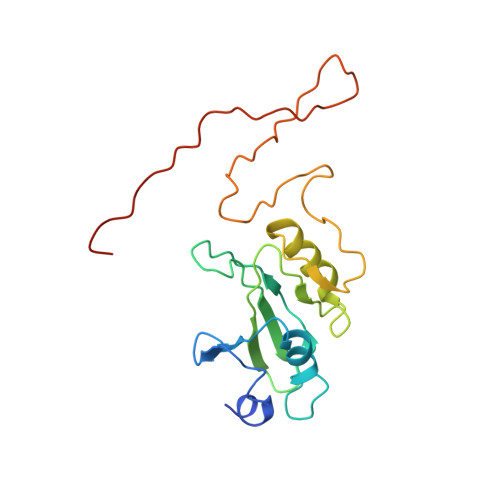
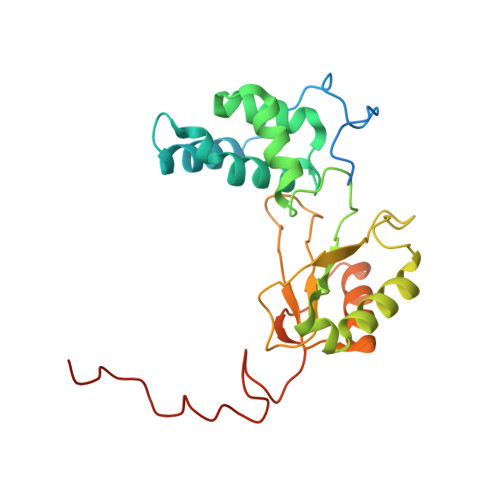
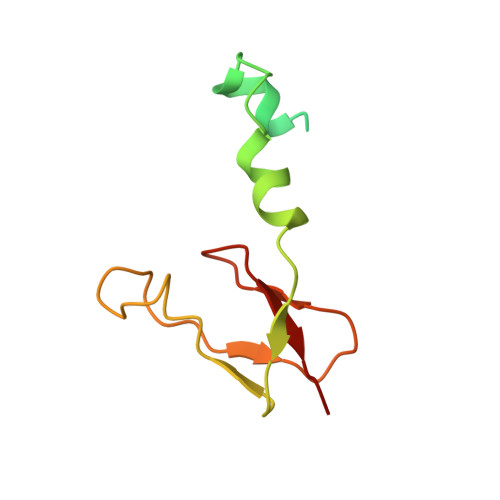
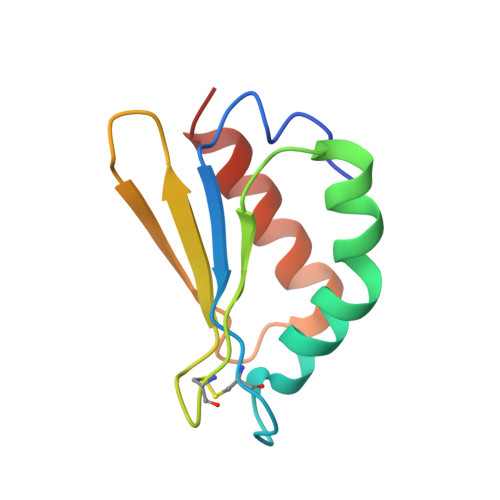
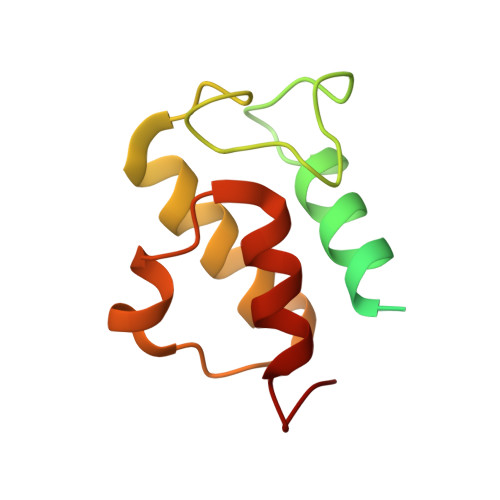
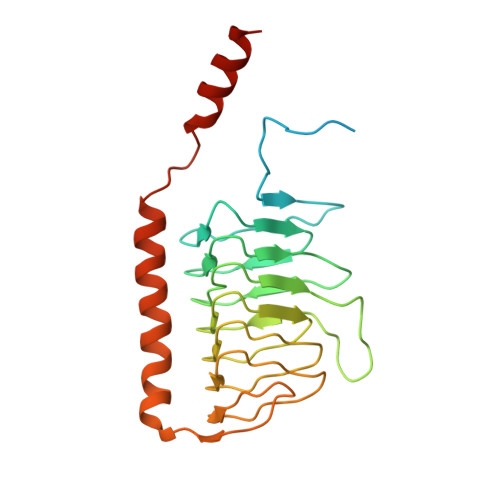
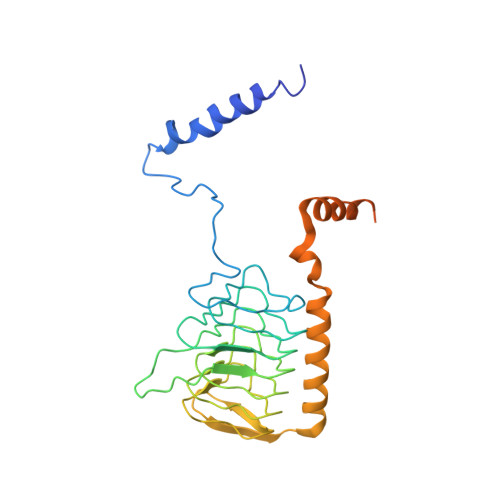
Specific features and assembly of the plant mitochondrial complex I revealed by cryo-EM.
Soufari, H., Parrot, C., Kuhn, L., Waltz, F., Hashem, Y.(2020) Nat Commun 11: 5195-5195
- PubMed: 33060577 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-18814-w
- Primary Citation Related Structures:
7A23, 7A24 - PubMed Abstract:
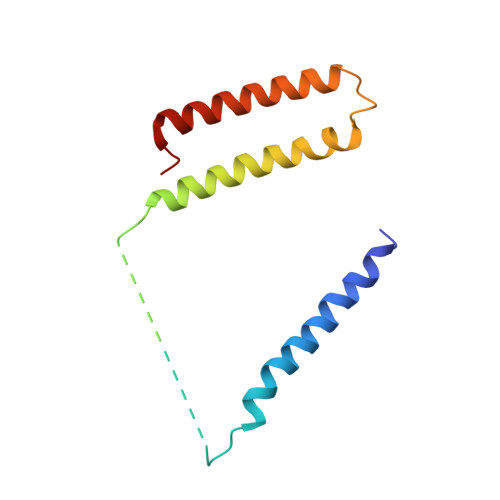
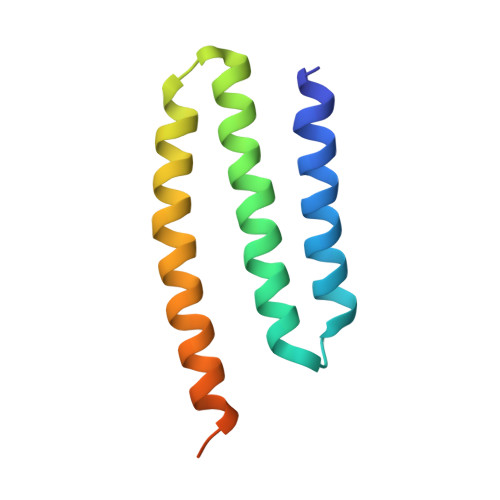
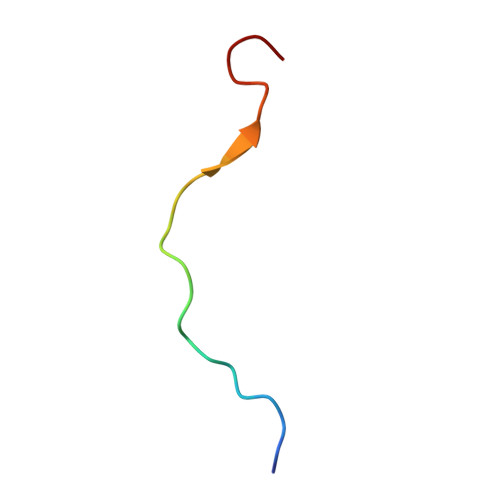
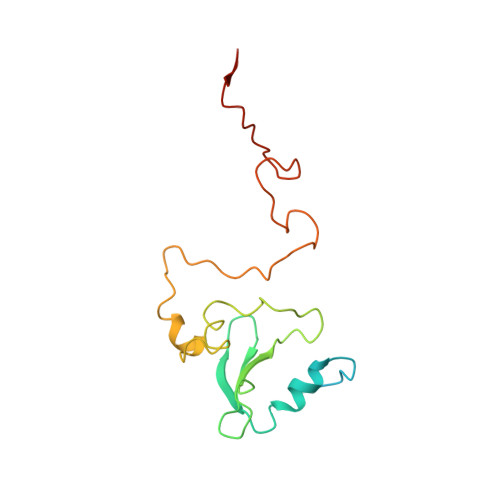
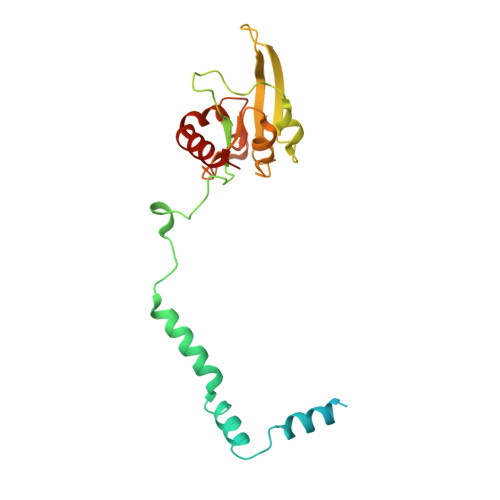
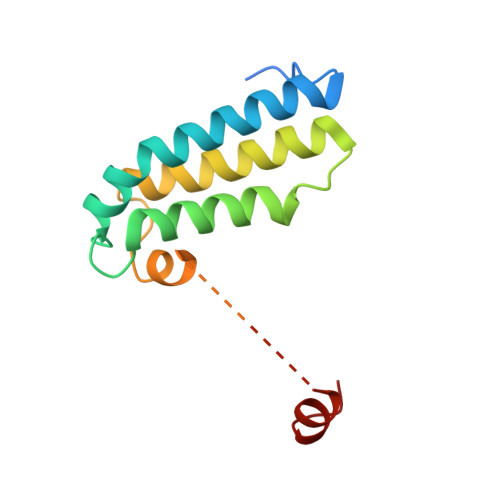
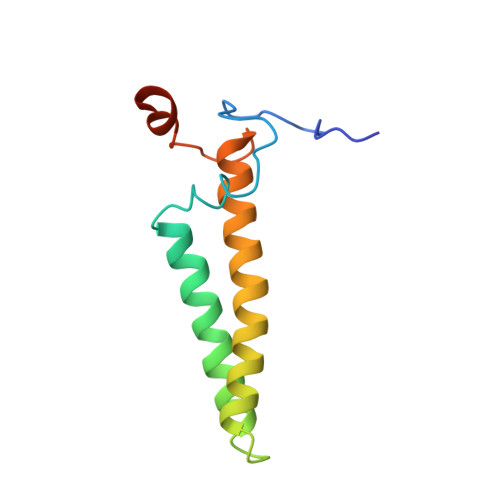
Mitochondria are the powerhouses of eukaryotic cells and the site of essential metabolic reactions. Complex I or NADH:ubiquinone oxidoreductase is the main entry site for electrons into the mitochondrial respiratory chain and constitutes the largest of the respiratory complexes. Its structure and composition vary across eukaryote species. However, high resolution structures are available only for one group of eukaryotes, opisthokonts. In plants, only biochemical studies were carried out, already hinting at the peculiar composition of complex I in the green lineage. Here, we report several cryo-electron microscopy structures of the plant mitochondrial complex I. We describe the structure and composition of the plant respiratory complex I, including the ancestral mitochondrial domain composed of the carbonic anhydrase. We show that the carbonic anhydrase is a heterotrimeric complex with only one conserved active site. This domain is crucial for the overall stability of complex I as well as a peculiar lipid complex composed of cardiolipin and phosphatidylinositols. Moreover, we also describe the structure of one of the plant-specific complex I assembly intermediates, lacking the whole P D module, in presence of the maturation factor GLDH. GLDH prevents the binding of the plant specific P1 protein, responsible for the linkage of the P P to the P D module.
- Institut Européen de Chimie et Biologie, U1212 Inserm, Université de Bordeaux, 2 rue R. Escarpit, F-33600, Pessac, France.
Organizational Affiliation: